U-5R
Derivation of equilibrium thermodynamic equations for U-5R system: isomerization in the binding-incompetent state of the receptor (many states)
2. Basic equilibrium equations
3. Derivation of equations for equilibrium concentrations
4. Prepare equations for a numeric solution
5. Save results on disk for future use
In this document, I am developing equilibrium thermodynamic equations to calculate concentrations of species at all points in titrations
clean up workspace
reset()
Set path to save results into:
ProjectName:="U-5R";
CurrentPath:="/Users/kovrigin_laptop/Documents/Workspace/Global_Analysis/IDAP/Mathematical_models/Equilibrium_thermodynamic_models/U-multi-path-models/nR/U-5R";
![]()
![]()
Binding and isomerization constants
All binding constants I am using are association constants.
These relationships serve as restraints for solve(), but not restrict these values in calculations!
K_A
K_A ;
assumeAlso(K_A > 0):
assumeAlso(K_A , R_)
![]()
To denote "starred" species, I will use s1 for *, s2 for **, etc.
K_B_s_1
K_B_s_1 ;
assumeAlso(K_B_s_1 > 0):
assumeAlso(K_B_s_1 , R_):
![]()
K_B_s_2
K_B_s_2 ;
assumeAlso(K_B_s_2 > 0):
assumeAlso(K_B_s_2 , R_):
![]()
K_B_s_3
K_B_s_3 ;
assumeAlso(K_B_s_3 > 0):
assumeAlso(K_B_s_3 , R_):
![]()
K_B_s_4
K_B_s_4 ;
assumeAlso(K_B_s_4 > 0):
assumeAlso(K_B_s_4 , R_):
![]()
K_B_s_5
K_B_s_5 ;
assumeAlso(K_B_s_5 > 0):
assumeAlso(K_B_s_5 , R_):
![]()
Total concentrations
Rtot - total concentration of the receptor
Rtot;
assumeAlso(Rtot>0):
assumeAlso(Rtot,R_):
![]()
Ltot - total concentration of the ligand
Ltot;
assumeAlso(Ltot>0):
assumeAlso(Ltot,R_):
![]()
Equilibrium concentrations
Req - equilibrium concentration of the binding-competent receptor form
Req;
assumeAlso(Req>0):
assumeAlso(Req<=Rtot):
assumeAlso(Req,R_):
![]()
Equilibrium concentrations of the binding incompetent receptor isomers Rn*
R_s_1eq
R_s_1eq;
assumeAlso(R_s_1eq>0):
assumeAlso(R_s_1eq<=Rtot):
assumeAlso(R_s_1eq,R_):
![]()
R_s_2eq
R_s_2eq;
assumeAlso(R_s_2eq>0):
assumeAlso(R_s_2eq<=Rtot):
assumeAlso(R_s_2eq,R_):
![]()
R_s_3eq
R_s_3eq;
assumeAlso(R_s_3eq>0):
assumeAlso(R_s_3eq<=Rtot):
assumeAlso(R_s_3eq,R_):
![]()
R_s_4eq
R_s_4eq;
assumeAlso(R_s_4eq>0):
assumeAlso(R_s_4eq<=Rtot):
assumeAlso(R_s_4eq,R_):
![]()
R_s_5eq
R_s_5eq;
assumeAlso(R_s_5eq>0):
assumeAlso(R_s_5eq<=Rtot):
assumeAlso(R_s_5eq,R_):
![]()
Equilibrium concentrations of other species
Leq - equilibrium concentration of a free ligand
Leq;
assumeAlso(Leq>0):
assumeAlso(Leq<Ltot):
assumeAlso(Leq,R_):
![]()
RLeq - equilibrium concentration of the receptor-ligand complex
RLeq;
assumeAlso(RLeq>0):
assumeAlso(RLeq<Rtot):
assumeAlso(RLeq<Ltot):
assumeAlso(RLeq,R_):
![]()
Check what we defined
anames(Properties,User);
![]()
2. Basic equilibrium equations
Mass conservation equations
eq2_1:= Rtot = Req + R_s_1eq + R_s_2eq + R_s_3eq + R_s_4eq + R_s_5eq + RLeq;
eq2_2:= Ltot = Leq + RLeq;
![]()
![]()
Equilibrium constants
eq2_3:= K_A = RLeq / (Req*Leq);

HINT: When increasing number of species: do not increase equation number, add a, b, c, ... modifier instead
because then you do NOT need to modify equation numbers in most of the following derivation!
eq2_4:= K_B_s_1 = R_s_1eq/Req
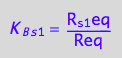
eq2_5:= K_B_s_2 = R_s_2eq/Req
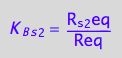
eq2_6:= K_B_s_3 = R_s_3eq/Req
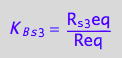
eq2_7:= K_B_s_4 = R_s_4eq/Req
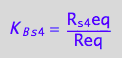
eq2_8:= K_B_s_5 = R_s_5eq/Req
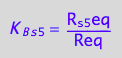
3. Derivation of equations for equilibrium concentrations
Express Leq as a function of all constants and total concentrations. If insoluble ---express Rtot=f(Leq and all constants).
Express the highest-order bound species first:
RL
eq2_3;
solve(%,RLeq):
%[1][1]:
eq3_1:= RLeq=%

![]()
Substitute into the mass conservation laws:
eq2_1;
% | eq3_1:
eq3_2:= %
![]()
![]()
eq2_2;
% | eq3_1:
eq3_3:= %
![]()
![]()
Express isomer concentrations
R*
eq2_4;
solve(%,R_s_1eq):
%[2][1]:
eq3_7:= R_s_1eq = %
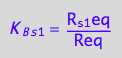
![]()
-> conservation laws
eq3_2;
% | eq3_7:
eq3_8:= %
![]()
![]()
R**
eq2_5;
solve(%, R_s_2eq):
%[2][1]:
eq3_9:= R_s_2eq = %
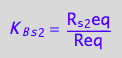
![]()
-> conservation laws
eq3_8;
% | eq3_9:
eq3_10:= %
![]()
![]()
R***
eq2_6;
solve(%, R_s_3eq):
%[2][1]:
eq3_11:= R_s_3eq = %
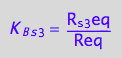
![]()
-> conservation laws
eq3_10;
% | eq3_11:
eq3_12:= %
![]()
![]()
R****
eq2_7;
solve(%, R_s_4eq):
%[2][1]:
eq3_13:= R_s_4eq = %
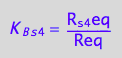
![]()
-> conservation laws
eq3_12;
% | eq3_13:
eq3_14:= %
![]()
![]()
R*****
eq2_8;
solve(%, R_s_5eq):
%[2][1]:
eq3_15:= R_s_5eq = %
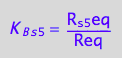
![]()
-> conservation laws
eq3_14;
% | eq3_15:
eq3_16:= %
![]()
![]()
Aim to obtain Rtot=f(Leq, constants) function
express Req from conservation law for ligand (Ltot=...)
eq3_3;
solve(%,Req):
%[1][1]:
eq3_17:= Req = %
![]()
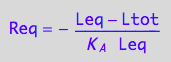
substitute in the conservation law for receptor: (Rtot=...)
eq3_16;
% | eq3_17;
temp1:=%[2];
temp2:=Simplify(%);
// test
temp1=temp2;
Simplify(%);
// Assemble a final equation
eq3_18:= Rtot =temp2
![]()


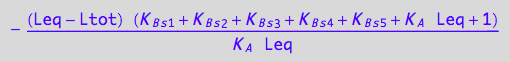

![]()
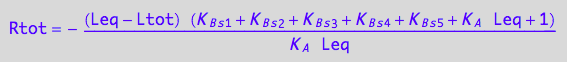
Attempt to solve for Leq
solution3_18:=solve(eq3_18, Leq)
![piecewise([-(K_B_s_1 + K_B_s_2 + K_B_s_3 + K_B_s_4 + K_B_s_5 + (K_A^2*Ltot^2 - 2*K_A^2*Ltot*Rtot + K_A^2*Rtot^2 + 2*K_A*K_B_s_1*Rtot + 2*K_A*K_B_s_2*Rtot + 2*K_A*K_B_s_3*Rtot + 2*K_A*K_B_s_4*Rtot + 2*K_A*K_B_s_5*Rtot + 2*K_A*Rtot + K_B_s_1^2 + 2*K_B_s_1*K_B_s_2 + 2*K_B_s_1*K_B_s_3 + 2*K_B_s_1*K_B_s_4 + 2*K_B_s_1*K_B_s_5 + 2*K_B_s_1 + K_B_s_2^2 + 2*K_B_s_2*K_B_s_3 + 2*K_B_s_2*K_B_s_4 + 2*K_B_s_2*K_B_s_5 + 2*K_B_s_2 + K_B_s_3^2 + 2*K_B_s_3*K_B_s_4 + 2*K_B_s_3*K_B_s_5 + 2*K_B_s_3 + K_B_s_4^2 + 2*K_B_s_4*K_B_s_5 + 2*K_B_s_4 + K_B_s_5^2 + 2*K_B_s_5 + 1)^(1/2) - K_A*Ltot + K_A*Rtot + 1)/(2*K_A) < Ltot and (K_A^2*Ltot^2 - 2*K_A^2*Ltot*Rtot + K_A^2*Rtot^2 + 2*K_A*K_B_s_1*Ltot + 2*K_A*K_B_s_1*Rtot + 2*K_A*K_B_s_2*Ltot + 2*K_A*K_B_s_2*Rtot + 2*K_A*K_B_s_3*Ltot + 2*K_A*K_B_s_3*Rtot + 2*K_A*K_B_s_4*Ltot + 2*K_A*K_B_s_4*Rtot + 2*K_A*K_B_s_5*Ltot + 2*K_A*K_B_s_5*Rtot + 2*K_A*Ltot + 2*K_A*Rtot + K_B_s_1^2 + 2*K_B_s_1*K_B_s_2 + 2*K_B_s_1*K_B_s_3 + 2*K_B_s_1*K_B_s_4 + 2*K_B_s_1*K_B_s_5 + 2*K_B_s_1 + K_B_s_2^2 + 2*K_B_s_2*K_B_s_3 + 2*K_B_s_2*K_B_s_4 + 2*K_B_s_2*K_B_s_5 + 2*K_B_s_2 + K_B_s_3^2 + 2*K_B_s_3*K_B_s_4 + 2*K_B_s_3*K_B_s_5 + 2*K_B_s_3 + K_B_s_4^2 + 2*K_B_s_4*K_B_s_5 + 2*K_B_s_4 + K_B_s_5^2 + 2*K_B_s_5 + 1)^(1/2) + K_A*Ltot = K_B_s_1 + K_B_s_2 + K_B_s_3 + K_B_s_4 + K_B_s_5 + K_A*Rtot + 1 and 2*K_A^2*Ltot*Rtot <= K_A^2*Ltot^2 + K_A^2*Rtot^2 + 2*K_A*K_B_s_1*Rtot + 2*K_A*K_B_s_2*Rtot + 2*K_A*K_B_s_3*Rtot + 2*K_A*K_B_s_4*Rtot + 2*K_A*K_B_s_5*Rtot + 2*K_A*Rtot + K_B_s_1^2 + 2*K_B_s_1*K_B_s_2 + 2*K_B_s_1*K_B_s_3 + 2*K_B_s_1*K_B_s_4 + 2*K_B_s_1*K_B_s_5 + 2*K_B_s_1 + K_B_s_2^2 + 2*K_B_s_2*K_B_s_3 + 2*K_B_s_2*K_B_s_4 + 2*K_B_s_2*K_B_s_5 + 2*K_B_s_2 + K_B_s_3^2 + 2*K_B_s_3*K_B_s_4 + 2*K_B_s_3*K_B_s_5 + 2*K_B_s_3 + K_B_s_4^2 + 2*K_B_s_4*K_B_s_5 + 2*K_B_s_4 + K_B_s_5^2 + 2*K_B_s_5 + 1 and K_B_s_1 + K_B_s_2 + K_B_s_3 + K_B_s_4 + K_B_s_5 + (K_A^2*Ltot^2 - 2*K_A^2*Ltot*Rtot + K_A^2*Rtot^2 + 2*K_A*K_B_s_1*Rtot + 2*K_A*K_B_s_2*Rtot + 2*K_A*K_B_s_3*Rtot + 2*K_A*K_B_s_4*Rtot + 2*K_A*K_B_s_5*Rtot + 2*K_A*Rtot + K_B_s_1^2 + 2*K_B_s_1*K_B_s_2 + 2*K_B_s_1*K_B_s_3 + 2*K_B_s_1*K_B_s_4 + 2*K_B_s_1*K_B_s_5 + 2*K_B_s_1 + K_B_s_2^2 + 2*K_B_s_2*K_B_s_3 + 2*K_B_s_2*K_B_s_4 + 2*K_B_s_2*K_B_s_5 + 2*K_B_s_2 + K_B_s_3^2 + 2*K_B_s_3*K_B_s_4 + 2*K_B_s_3*K_B_s_5 + 2*K_B_s_3 + K_B_s_4^2 + 2*K_B_s_4*K_B_s_5 + 2*K_B_s_4 + K_B_s_5^2 + 2*K_B_s_5 + 1)^(1/2) + K_A*Rtot + 1 < K_A*Ltot and K_B_s_1 + K_B_s_2 + K_B_s_3 + K_B_s_4 + K_B_s_5 + (K_A^2*Ltot^2 - 2*K_A^2*Ltot*Rtot + K_A^2*Rtot^2 + 2*K_A*K_B_s_1*Ltot + 2*K_A*K_B_s_1*Rtot + 2*K_A*K_B_s_2*Ltot + 2*K_A*K_B_s_2*Rtot + 2*K_A*K_B_s_3*Ltot + 2*K_A*K_B_s_3*Rtot + 2*K_A*K_B_s_4*Ltot + 2*K_A*K_B_s_4*Rtot + 2*K_A*K_B_s_5*Ltot + 2*K_A*K_B_s_5*Rtot + 2*K_A*Ltot + 2*K_A*Rtot + K_B_s_1^2 + 2*K_B_s_1*K_B_s_2 + 2*K_B_s_1*K_B_s_3 + 2*K_B_s_1*K_B_s_4 + 2*K_B_s_1*K_B_s_5 + 2*K_B_s_1 + K_B_s_2^2 + 2*K_B_s_2*K_B_s_3 + 2*K_B_s_2*K_B_s_4 + 2*K_B_s_2*K_B_s_5 + 2*K_B_s_2 + K_B_s_3^2 + 2*K_B_s_3*K_B_s_4 + 2*K_B_s_3*K_B_s_5 + 2*K_B_s_3 + K_B_s_4^2 + 2*K_B_s_4*K_B_s_5 + 2*K_B_s_4 + K_B_s_5^2 + 2*K_B_s_5 + 1)^(1/2) + K_A*Rtot + 1 <> K_A*Ltot, {-(K_B_s_1 + K_B_s_2 + K_B_s_3 + K_B_s_4 + K_B_s_5 + (K_A^2*Ltot^2 - 2*K_A^2*Ltot*Rtot + K_A^2*Rtot^2 + 2*K_A*K_B_s_1*Rtot + 2*K_A*K_B_s_2*Rtot + 2*K_A*K_B_s_3*Rtot + 2*K_A*K_B_s_4*Rtot + 2*K_A*K_B_s_5*Rtot + 2*K_A*Rtot + K_B_s_1^2 + 2*K_B_s_1*K_B_s_2 + 2*K_B_s_1*K_B_s_3 + 2*K_B_s_1*K_B_s_4 + 2*K_B_s_1*K_B_s_5 + 2*K_B_s_1 + K_B_s_2^2 + 2*K_B_s_2*K_B_s_3 + 2*K_B_s_2*K_B_s_4 + 2*K_B_s_2*K_B_s_5 + 2*K_B_s_2 + K_B_s_3^2 + 2*K_B_s_3*K_B_s_4 + 2*K_B_s_3*K_B_s_5 + 2*K_B_s_3 + K_B_s_4^2 + 2*K_B_s_4*K_B_s_5 + 2*K_B_s_4 + K_B_s_5^2 + 2*K_B_s_5 + 1)^(1/2) - K_A*Ltot + K_A*Rtot + 1)/(2*K_A)}], [K_B_s_1 + K_B_s_2 + K_B_s_3 + K_B_s_4 + K_B_s_5 - (K_A^2*Ltot^2 - 2*K_A^2*Ltot*Rtot + K_A^2*Rtot^2 + 2*K_A*K_B_s_1*Rtot + 2*K_A*K_B_s_2*Rtot + 2*K_A*K_B_s_3*Rtot + 2*K_A*K_B_s_4*Rtot + 2*K_A*K_B_s_5*Rtot + 2*K_A*Rtot + K_B_s_1^2 + 2*K_B_s_1*K_B_s_2 + 2*K_B_s_1*K_B_s_3 + 2*K_B_s_1*K_B_s_4 + 2*K_B_s_1*K_B_s_5 + 2*K_B_s_1 + K_B_s_2^2 + 2*K_B_s_2*K_B_s_3 + 2*K_B_s_2*K_B_s_4 + 2*K_B_s_2*K_B_s_5 + 2*K_B_s_2 + K_B_s_3^2 + 2*K_B_s_3*K_B_s_4 + 2*K_B_s_3*K_B_s_5 + 2*K_B_s_3 + K_B_s_4^2 + 2*K_B_s_4*K_B_s_5 + 2*K_B_s_4 + K_B_s_5^2 + 2*K_B_s_5 + 1)^(1/2) + K_A*Rtot + 1 < K_A*Ltot and (K_A^2*Ltot^2 - 2*K_A^2*Ltot*Rtot + K_A^2*Rtot^2 + 2*K_A*K_B_s_1*Ltot + 2*K_A*K_B_s_1*Rtot + 2*K_A*K_B_s_2*Ltot + 2*K_A*K_B_s_2*Rtot + 2*K_A*K_B_s_3*Ltot + 2*K_A*K_B_s_3*Rtot + 2*K_A*K_B_s_4*Ltot + 2*K_A*K_B_s_4*Rtot + 2*K_A*K_B_s_5*Ltot + 2*K_A*K_B_s_5*Rtot + 2*K_A*Ltot + 2*K_A*Rtot + K_B_s_1^2 + 2*K_B_s_1*K_B_s_2 + 2*K_B_s_1*K_B_s_3 + 2*K_B_s_1*K_B_s_4 + 2*K_B_s_1*K_B_s_5 + 2*K_B_s_1 + K_B_s_2^2 + 2*K_B_s_2*K_B_s_3 + 2*K_B_s_2*K_B_s_4 + 2*K_B_s_2*K_B_s_5 + 2*K_B_s_2 + K_B_s_3^2 + 2*K_B_s_3*K_B_s_4 + 2*K_B_s_3*K_B_s_5 + 2*K_B_s_3 + K_B_s_4^2 + 2*K_B_s_4*K_B_s_5 + 2*K_B_s_4 + K_B_s_5^2 + 2*K_B_s_5 + 1)^(1/2) + K_A*Ltot <> K_B_s_1 + K_B_s_2 + K_B_s_3 + K_B_s_4 + K_B_s_5 + K_A*Rtot + 1 and -(K_B_s_1 + K_B_s_2 + K_B_s_3 + K_B_s_4 + K_B_s_5 - (K_A^2*Ltot^2 - 2*K_A^2*Ltot*Rtot + K_A^2*Rtot^2 + 2*K_A*K_B_s_1*Rtot + 2*K_A*K_B_s_2*Rtot + 2*K_A*K_B_s_3*Rtot + 2*K_A*K_B_s_4*Rtot + 2*K_A*K_B_s_5*Rtot + 2*K_A*Rtot + K_B_s_1^2 + 2*K_B_s_1*K_B_s_2 + 2*K_B_s_1*K_B_s_3 + 2*K_B_s_1*K_B_s_4 + 2*K_B_s_1*K_B_s_5 + 2*K_B_s_1 + K_B_s_2^2 + 2*K_B_s_2*K_B_s_3 + 2*K_B_s_2*K_B_s_4 + 2*K_B_s_2*K_B_s_5 + 2*K_B_s_2 + K_B_s_3^2 + 2*K_B_s_3*K_B_s_4 + 2*K_B_s_3*K_B_s_5 + 2*K_B_s_3 + K_B_s_4^2 + 2*K_B_s_4*K_B_s_5 + 2*K_B_s_4 + K_B_s_5^2 + 2*K_B_s_5 + 1)^(1/2) - K_A*Ltot + K_A*Rtot + 1)/(2*K_A) < Ltot and 2*K_A^2*Ltot*Rtot <= K_A^2*Ltot^2 + K_A^2*Rtot^2 + 2*K_A*K_B_s_1*Rtot + 2*K_A*K_B_s_2*Rtot + 2*K_A*K_B_s_3*Rtot + 2*K_A*K_B_s_4*Rtot + 2*K_A*K_B_s_5*Rtot + 2*K_A*Rtot + K_B_s_1^2 + 2*K_B_s_1*K_B_s_2 + 2*K_B_s_1*K_B_s_3 + 2*K_B_s_1*K_B_s_4 + 2*K_B_s_1*K_B_s_5 + 2*K_B_s_1 + K_B_s_2^2 + 2*K_B_s_2*K_B_s_3 + 2*K_B_s_2*K_B_s_4 + 2*K_B_s_2*K_B_s_5 + 2*K_B_s_2 + K_B_s_3^2 + 2*K_B_s_3*K_B_s_4 + 2*K_B_s_3*K_B_s_5 + 2*K_B_s_3 + K_B_s_4^2 + 2*K_B_s_4*K_B_s_5 + 2*K_B_s_4 + K_B_s_5^2 + 2*K_B_s_5 + 1 and K_B_s_1 + K_B_s_2 + K_B_s_3 + K_B_s_4 + K_B_s_5 + (K_A^2*Ltot^2 - 2*K_A^2*Ltot*Rtot + K_A^2*Rtot^2 + 2*K_A*K_B_s_1*Ltot + 2*K_A*K_B_s_1*Rtot + 2*K_A*K_B_s_2*Ltot + 2*K_A*K_B_s_2*Rtot + 2*K_A*K_B_s_3*Ltot + 2*K_A*K_B_s_3*Rtot + 2*K_A*K_B_s_4*Ltot + 2*K_A*K_B_s_4*Rtot + 2*K_A*K_B_s_5*Ltot + 2*K_A*K_B_s_5*Rtot + 2*K_A*Ltot + 2*K_A*Rtot + K_B_s_1^2 + 2*K_B_s_1*K_B_s_2 + 2*K_B_s_1*K_B_s_3 + 2*K_B_s_1*K_B_s_4 + 2*K_B_s_1*K_B_s_5 + 2*K_B_s_1 + K_B_s_2^2 + 2*K_B_s_2*K_B_s_3 + 2*K_B_s_2*K_B_s_4 + 2*K_B_s_2*K_B_s_5 + 2*K_B_s_2 + K_B_s_3^2 + 2*K_B_s_3*K_B_s_4 + 2*K_B_s_3*K_B_s_5 + 2*K_B_s_3 + K_B_s_4^2 + 2*K_B_s_4*K_B_s_5 + 2*K_B_s_4 + K_B_s_5^2 + 2*K_B_s_5 + 1)^(1/2) + K_A*Rtot + 1 = K_A*Ltot, {-(K_B_s_1 + K_B_s_2 + K_B_s_3 + K_B_s_4 + K_B_s_5 - (K_A^2*Ltot^2 - 2*K_A^2*Ltot*Rtot + K_A^2*Rtot^2 + 2*K_A*K_B_s_1*Rtot + 2*K_A*K_B_s_2*Rtot + 2*K_A*K_B_s_3*Rtot + 2*K_A*K_B_s_4*Rtot + 2*K_A*K_B_s_5*Rtot + 2*K_A*Rtot + K_B_s_1^2 + 2*K_B_s_1*K_B_s_2 + 2*K_B_s_1*K_B_s_3 + 2*K_B_s_1*K_B_s_4 + 2*K_B_s_1*K_B_s_5 + 2*K_B_s_1 + K_B_s_2^2 + 2*K_B_s_2*K_B_s_3 + 2*K_B_s_2*K_B_s_4 + 2*K_B_s_2*K_B_s_5 + 2*K_B_s_2 + K_B_s_3^2 + 2*K_B_s_3*K_B_s_4 + 2*K_B_s_3*K_B_s_5 + 2*K_B_s_3 + K_B_s_4^2 + 2*K_B_s_4*K_B_s_5 + 2*K_B_s_4 + K_B_s_5^2 + 2*K_B_s_5 + 1)^(1/2) - K_A*Ltot + K_A*Rtot + 1)/(2*K_A)}], [-(K_B_s_1 + K_B_s_2 + K_B_s_3 + K_B_s_4 + K_B_s_5 + (K_A^2*Ltot^2 - 2*K_A^2*Ltot*Rtot + K_A^2*Rtot^2 + 2*K_A*K_B_s_1*Ltot + 2*K_A*K_B_s_1*Rtot + 2*K_A*K_B_s_2*Ltot + 2*K_A*K_B_s_2*Rtot + 2*K_A*K_B_s_3*Ltot + 2*K_A*K_B_s_3*Rtot + 2*K_A*K_B_s_4*Ltot + 2*K_A*K_B_s_4*Rtot + 2*K_A*K_B_s_5*Ltot + 2*K_A*K_B_s_5*Rtot + 2*K_A*Ltot + 2*K_A*Rtot + K_B_s_1^2 + 2*K_B_s_1*K_B_s_2 + 2*K_B_s_1*K_B_s_3 + 2*K_B_s_1*K_B_s_4 + 2*K_B_s_1*K_B_s_5 + 2*K_B_s_1 + K_B_s_2^2 + 2*K_B_s_2*K_B_s_3 + 2*K_B_s_2*K_B_s_4 + 2*K_B_s_2*K_B_s_5 + 2*K_B_s_2 + K_B_s_3^2 + 2*K_B_s_3*K_B_s_4 + 2*K_B_s_3*K_B_s_5 + 2*K_B_s_3 + K_B_s_4^2 + 2*K_B_s_4*K_B_s_5 + 2*K_B_s_4 + K_B_s_5^2 + 2*K_B_s_5 + 1)^(1/2) - K_A*Ltot + K_A*Rtot + 1)/(2*K_A) < Ltot and K_B_s_1 + K_B_s_2 + K_B_s_3 + K_B_s_4 + K_B_s_5 - (K_A^2*Ltot^2 - 2*K_A^2*Ltot*Rtot + K_A^2*Rtot^2 + 2*K_A*K_B_s_1*Ltot + 2*K_A*K_B_s_1*Rtot + 2*K_A*K_B_s_2*Ltot + 2*K_A*K_B_s_2*Rtot + 2*K_A*K_B_s_3*Ltot + 2*K_A*K_B_s_3*Rtot + 2*K_A*K_B_s_4*Ltot + 2*K_A*K_B_s_4*Rtot + 2*K_A*K_B_s_5*Ltot + 2*K_A*K_B_s_5*Rtot + 2*K_A*Ltot + 2*K_A*Rtot + K_B_s_1^2 + 2*K_B_s_1*K_B_s_2 + 2*K_B_s_1*K_B_s_3 + 2*K_B_s_1*K_B_s_4 + 2*K_B_s_1*K_B_s_5 + 2*K_B_s_1 + K_B_s_2^2 + 2*K_B_s_2*K_B_s_3 + 2*K_B_s_2*K_B_s_4 + 2*K_B_s_2*K_B_s_5 + 2*K_B_s_2 + K_B_s_3^2 + 2*K_B_s_3*K_B_s_4 + 2*K_B_s_3*K_B_s_5 + 2*K_B_s_3 + K_B_s_4^2 + 2*K_B_s_4*K_B_s_5 + 2*K_B_s_4 + K_B_s_5^2 + 2*K_B_s_5 + 1)^(1/2) + K_A*Rtot + 1 < K_A*Ltot and (K_A^2*Ltot^2 - 2*K_A^2*Ltot*Rtot + K_A^2*Rtot^2 + 2*K_A*K_B_s_1*Ltot + 2*K_A*K_B_s_1*Rtot + 2*K_A*K_B_s_2*Ltot + 2*K_A*K_B_s_2*Rtot + 2*K_A*K_B_s_3*Ltot + 2*K_A*K_B_s_3*Rtot + 2*K_A*K_B_s_4*Ltot + 2*K_A*K_B_s_4*Rtot + 2*K_A*K_B_s_5*Ltot + 2*K_A*K_B_s_5*Rtot + 2*K_A*Ltot + 2*K_A*Rtot + K_B_s_1^2 + 2*K_B_s_1*K_B_s_2 + 2*K_B_s_1*K_B_s_3 + 2*K_B_s_1*K_B_s_4 + 2*K_B_s_1*K_B_s_5 + 2*K_B_s_1 + K_B_s_2^2 + 2*K_B_s_2*K_B_s_3 + 2*K_B_s_2*K_B_s_4 + 2*K_B_s_2*K_B_s_5 + 2*K_B_s_2 + K_B_s_3^2 + 2*K_B_s_3*K_B_s_4 + 2*K_B_s_3*K_B_s_5 + 2*K_B_s_3 + K_B_s_4^2 + 2*K_B_s_4*K_B_s_5 + 2*K_B_s_4 + K_B_s_5^2 + 2*K_B_s_5 + 1)^(1/2) + K_A*Ltot <> K_B_s_1 + K_B_s_2 + K_B_s_3 + K_B_s_4 + K_B_s_5 + K_A*Rtot + 1 and -(K_B_s_1 + K_B_s_2 + K_B_s_3 + K_B_s_4 + K_B_s_5 - (K_A^2*Ltot^2 - 2*K_A^2*Ltot*Rtot + K_A^2*Rtot^2 + 2*K_A*K_B_s_1*Ltot + 2*K_A*K_B_s_1*Rtot + 2*K_A*K_B_s_2*Ltot + 2*K_A*K_B_s_2*Rtot + 2*K_A*K_B_s_3*Ltot + 2*K_A*K_B_s_3*Rtot + 2*K_A*K_B_s_4*Ltot + 2*K_A*K_B_s_4*Rtot + 2*K_A*K_B_s_5*Ltot + 2*K_A*K_B_s_5*Rtot + 2*K_A*Ltot + 2*K_A*Rtot + K_B_s_1^2 + 2*K_B_s_1*K_B_s_2 + 2*K_B_s_1*K_B_s_3 + 2*K_B_s_1*K_B_s_4 + 2*K_B_s_1*K_B_s_5 + 2*K_B_s_1 + K_B_s_2^2 + 2*K_B_s_2*K_B_s_3 + 2*K_B_s_2*K_B_s_4 + 2*K_B_s_2*K_B_s_5 + 2*K_B_s_2 + K_B_s_3^2 + 2*K_B_s_3*K_B_s_4 + 2*K_B_s_3*K_B_s_5 + 2*K_B_s_3 + K_B_s_4^2 + 2*K_B_s_4*K_B_s_5 + 2*K_B_s_4 + K_B_s_5^2 + 2*K_B_s_5 + 1)^(1/2) - K_A*Ltot + K_A*Rtot + 1)/(2*K_A) < Ltot and 2*K_A^2*Ltot*Rtot <= K_A^2*Ltot^2 + K_A^2*Rtot^2 + 2*K_A*K_B_s_1*Ltot + 2*K_A*K_B_s_1*Rtot + 2*K_A*K_B_s_2*Ltot + 2*K_A*K_B_s_2*Rtot + 2*K_A*K_B_s_3*Ltot + 2*K_A*K_B_s_3*Rtot + 2*K_A*K_B_s_4*Ltot + 2*K_A*K_B_s_4*Rtot + 2*K_A*K_B_s_5*Ltot + 2*K_A*K_B_s_5*Rtot + 2*K_A*Ltot + 2*K_A*Rtot + K_B_s_1^2 + 2*K_B_s_1*K_B_s_2 + 2*K_B_s_1*K_B_s_3 + 2*K_B_s_1*K_B_s_4 + 2*K_B_s_1*K_B_s_5 + 2*K_B_s_1 + K_B_s_2^2 + 2*K_B_s_2*K_B_s_3 + 2*K_B_s_2*K_B_s_4 + 2*K_B_s_2*K_B_s_5 + 2*K_B_s_2 + K_B_s_3^2 + 2*K_B_s_3*K_B_s_4 + 2*K_B_s_3*K_B_s_5 + 2*K_B_s_3 + K_B_s_4^2 + 2*K_B_s_4*K_B_s_5 + 2*K_B_s_4 + K_B_s_5^2 + 2*K_B_s_5 + 1 and K_B_s_1 + K_B_s_2 + K_B_s_3 + K_B_s_4 + K_B_s_5 + (K_A^2*Ltot^2 - 2*K_A^2*Ltot*Rtot + K_A^2*Rtot^2 + 2*K_A*K_B_s_1*Ltot + 2*K_A*K_B_s_1*Rtot + 2*K_A*K_B_s_2*Ltot + 2*K_A*K_B_s_2*Rtot + 2*K_A*K_B_s_3*Ltot + 2*K_A*K_B_s_3*Rtot + 2*K_A*K_B_s_4*Ltot + 2*K_A*K_B_s_4*Rtot + 2*K_A*K_B_s_5*Ltot + 2*K_A*K_B_s_5*Rtot + 2*K_A*Ltot + 2*K_A*Rtot + K_B_s_1^2 + 2*K_B_s_1*K_B_s_2 + 2*K_B_s_1*K_B_s_3 + 2*K_B_s_1*K_B_s_4 + 2*K_B_s_1*K_B_s_5 + 2*K_B_s_1 + K_B_s_2^2 + 2*K_B_s_2*K_B_s_3 + 2*K_B_s_2*K_B_s_4 + 2*K_B_s_2*K_B_s_5 + 2*K_B_s_2 + K_B_s_3^2 + 2*K_B_s_3*K_B_s_4 + 2*K_B_s_3*K_B_s_5 + 2*K_B_s_3 + K_B_s_4^2 + 2*K_B_s_4*K_B_s_5 + 2*K_B_s_4 + K_B_s_5^2 + 2*K_B_s_5 + 1)^(1/2) + K_A*Rtot + 1 < K_A*Ltot, {-(K_B_s_1 + K_B_s_2 + K_B_s_3 + K_B_s_4 + K_B_s_5 + (K_A^2*Ltot^2 - 2*K_A^2*Ltot*Rtot + K_A^2*Rtot^2 + 2*K_A*K_B_s_1*Ltot + 2*K_A*K_B_s_1*Rtot + 2*K_A*K_B_s_2*Ltot + 2*K_A*K_B_s_2*Rtot + 2*K_A*K_B_s_3*Ltot + 2*K_A*K_B_s_3*Rtot + 2*K_A*K_B_s_4*Ltot + 2*K_A*K_B_s_4*Rtot + 2*K_A*K_B_s_5*Ltot + 2*K_A*K_B_s_5*Rtot + 2*K_A*Ltot + 2*K_A*Rtot + K_B_s_1^2 + 2*K_B_s_1*K_B_s_2 + 2*K_B_s_1*K_B_s_3 + 2*K_B_s_1*K_B_s_4 + 2*K_B_s_1*K_B_s_5 + 2*K_B_s_1 + K_B_s_2^2 + 2*K_B_s_2*K_B_s_3 + 2*K_B_s_2*K_B_s_4 + 2*K_B_s_2*K_B_s_5 + 2*K_B_s_2 + K_B_s_3^2 + 2*K_B_s_3*K_B_s_4 + 2*K_B_s_3*K_B_s_5 + 2*K_B_s_3 + K_B_s_4^2 + 2*K_B_s_4*K_B_s_5 + 2*K_B_s_4 + K_B_s_5^2 + 2*K_B_s_5 + 1)^(1/2) - K_A*Ltot + K_A*Rtot + 1)/(2*K_A), -(K_B_s_1 + K_B_s_2 + K_B_s_3 + K_B_s_4 + K_B_s_5 - (K_A^2*Ltot^2 - 2*K_A^2*Ltot*Rtot + K_A^2*Rtot^2 + 2*K_A*K_B_s_1*Ltot + 2*K_A*K_B_s_1*Rtot + 2*K_A*K_B_s_2*Ltot + 2*K_A*K_B_s_2*Rtot + 2*K_A*K_B_s_3*Ltot + 2*K_A*K_B_s_3*Rtot + 2*K_A*K_B_s_4*Ltot + 2*K_A*K_B_s_4*Rtot + 2*K_A*K_B_s_5*Ltot + 2*K_A*K_B_s_5*Rtot + 2*K_A*Ltot + 2*K_A*Rtot + K_B_s_1^2 + 2*K_B_s_1*K_B_s_2 + 2*K_B_s_1*K_B_s_3 + 2*K_B_s_1*K_B_s_4 + 2*K_B_s_1*K_B_s_5 + 2*K_B_s_1 + K_B_s_2^2 + 2*K_B_s_2*K_B_s_3 + 2*K_B_s_2*K_B_s_4 + 2*K_B_s_2*K_B_s_5 + 2*K_B_s_2 + K_B_s_3^2 + 2*K_B_s_3*K_B_s_4 + 2*K_B_s_3*K_B_s_5 + 2*K_B_s_3 + K_B_s_4^2 + 2*K_B_s_4*K_B_s_5 + 2*K_B_s_4 + K_B_s_5^2 + 2*K_B_s_5 + 1)^(1/2) - K_A*Ltot + K_A*Rtot + 1)/(2*K_A)}], [(not K_B_s_1 + K_B_s_2 + K_B_s_3 + K_B_s_4 + K_B_s_5 - (K_A^2*Ltot^2 - 2*K_A^2*Ltot*Rtot + K_A^2*Rtot^2 + 2*K_A*K_B_s_1*Ltot + 2*K_A*K_B_s_1*Rtot + 2*K_A*K_B_s_2*Ltot + 2*K_A*K_B_s_2*Rtot + 2*K_A*K_B_s_3*Ltot + 2*K_A*K_B_s_3*Rtot + 2*K_A*K_B_s_4*Ltot + 2*K_A*K_B_s_4*Rtot + 2*K_A*K_B_s_5*Ltot + 2*K_A*K_B_s_5*Rtot + 2*K_A*Ltot + 2*K_A*Rtot + K_B_s_1^2 + 2*K_B_s_1*K_B_s_2 + 2*K_B_s_1*K_B_s_3 + 2*K_B_s_1*K_B_s_4 + 2*K_B_s_1*K_B_s_5 + 2*K_B_s_1 + K_B_s_2^2 + 2*K_B_s_2*K_B_s_3 + 2*K_B_s_2*K_B_s_4 + 2*K_B_s_2*K_B_s_5 + 2*K_B_s_2 + K_B_s_3^2 + 2*K_B_s_3*K_B_s_4 + 2*K_B_s_3*K_B_s_5 + 2*K_B_s_3 + K_B_s_4^2 + 2*K_B_s_4*K_B_s_5 + 2*K_B_s_4 + K_B_s_5^2 + 2*K_B_s_5 + 1)^(1/2) + K_A*Rtot + 1 < K_A*Ltot or not -(K_B_s_1 + K_B_s_2 + K_B_s_3 + K_B_s_4 + K_B_s_5 - (K_A^2*Ltot^2 - 2*K_A^2*Ltot*Rtot + K_A^2*Rtot^2 + 2*K_A*K_B_s_1*Ltot + 2*K_A*K_B_s_1*Rtot + 2*K_A*K_B_s_2*Ltot + 2*K_A*K_B_s_2*Rtot + 2*K_A*K_B_s_3*Ltot + 2*K_A*K_B_s_3*Rtot + 2*K_A*K_B_s_4*Ltot + 2*K_A*K_B_s_4*Rtot + 2*K_A*K_B_s_5*Ltot + 2*K_A*K_B_s_5*Rtot + 2*K_A*Ltot + 2*K_A*Rtot + K_B_s_1^2 + 2*K_B_s_1*K_B_s_2 + 2*K_B_s_1*K_B_s_3 + 2*K_B_s_1*K_B_s_4 + 2*K_B_s_1*K_B_s_5 + 2*K_B_s_1 + K_B_s_2^2 + 2*K_B_s_2*K_B_s_3 + 2*K_B_s_2*K_B_s_4 + 2*K_B_s_2*K_B_s_5 + 2*K_B_s_2 + K_B_s_3^2 + 2*K_B_s_3*K_B_s_4 + 2*K_B_s_3*K_B_s_5 + 2*K_B_s_3 + K_B_s_4^2 + 2*K_B_s_4*K_B_s_5 + 2*K_B_s_4 + K_B_s_5^2 + 2*K_B_s_5 + 1)^(1/2) - K_A*Ltot + K_A*Rtot + 1)/(2*K_A) < Ltot) and -(K_B_s_1 + K_B_s_2 + K_B_s_3 + K_B_s_4 + K_B_s_5 + (K_A^2*Ltot^2 - 2*K_A^2*Ltot*Rtot + K_A^2*Rtot^2 + 2*K_A*K_B_s_1*Ltot + 2*K_A*K_B_s_1*Rtot + 2*K_A*K_B_s_2*Ltot + 2*K_A*K_B_s_2*Rtot + 2*K_A*K_B_s_3*Ltot + 2*K_A*K_B_s_3*Rtot + 2*K_A*K_B_s_4*Ltot + 2*K_A*K_B_s_4*Rtot + 2*K_A*K_B_s_5*Ltot + 2*K_A*K_B_s_5*Rtot + 2*K_A*Ltot + 2*K_A*Rtot + K_B_s_1^2 + 2*K_B_s_1*K_B_s_2 + 2*K_B_s_1*K_B_s_3 + 2*K_B_s_1*K_B_s_4 + 2*K_B_s_1*K_B_s_5 + 2*K_B_s_1 + K_B_s_2^2 + 2*K_B_s_2*K_B_s_3 + 2*K_B_s_2*K_B_s_4 + 2*K_B_s_2*K_B_s_5 + 2*K_B_s_2 + K_B_s_3^2 + 2*K_B_s_3*K_B_s_4 + 2*K_B_s_3*K_B_s_5 + 2*K_B_s_3 + K_B_s_4^2 + 2*K_B_s_4*K_B_s_5 + 2*K_B_s_4 + K_B_s_5^2 + 2*K_B_s_5 + 1)^(1/2) - K_A*Ltot + K_A*Rtot + 1)/(2*K_A) < Ltot and (K_A^2*Ltot^2 - 2*K_A^2*Ltot*Rtot + K_A^2*Rtot^2 + 2*K_A*K_B_s_1*Ltot + 2*K_A*K_B_s_1*Rtot + 2*K_A*K_B_s_2*Ltot + 2*K_A*K_B_s_2*Rtot + 2*K_A*K_B_s_3*Ltot + 2*K_A*K_B_s_3*Rtot + 2*K_A*K_B_s_4*Ltot + 2*K_A*K_B_s_4*Rtot + 2*K_A*K_B_s_5*Ltot + 2*K_A*K_B_s_5*Rtot + 2*K_A*Ltot + 2*K_A*Rtot + K_B_s_1^2 + 2*K_B_s_1*K_B_s_2 + 2*K_B_s_1*K_B_s_3 + 2*K_B_s_1*K_B_s_4 + 2*K_B_s_1*K_B_s_5 + 2*K_B_s_1 + K_B_s_2^2 + 2*K_B_s_2*K_B_s_3 + 2*K_B_s_2*K_B_s_4 + 2*K_B_s_2*K_B_s_5 + 2*K_B_s_2 + K_B_s_3^2 + 2*K_B_s_3*K_B_s_4 + 2*K_B_s_3*K_B_s_5 + 2*K_B_s_3 + K_B_s_4^2 + 2*K_B_s_4*K_B_s_5 + 2*K_B_s_4 + K_B_s_5^2 + 2*K_B_s_5 + 1)^(1/2) + K_A*Ltot <> K_B_s_1 + K_B_s_2 + K_B_s_3 + K_B_s_4 + K_B_s_5 + K_A*Rtot + 1 and 2*K_A^2*Ltot*Rtot <= K_A^2*Ltot^2 + K_A^2*Rtot^2 + 2*K_A*K_B_s_1*Ltot + 2*K_A*K_B_s_1*Rtot + 2*K_A*K_B_s_2*Ltot + 2*K_A*K_B_s_2*Rtot + 2*K_A*K_B_s_3*Ltot + 2*K_A*K_B_s_3*Rtot + 2*K_A*K_B_s_4*Ltot + 2*K_A*K_B_s_4*Rtot + 2*K_A*K_B_s_5*Ltot + 2*K_A*K_B_s_5*Rtot + 2*K_A*Ltot + 2*K_A*Rtot + K_B_s_1^2 + 2*K_B_s_1*K_B_s_2 + 2*K_B_s_1*K_B_s_3 + 2*K_B_s_1*K_B_s_4 + 2*K_B_s_1*K_B_s_5 + 2*K_B_s_1 + K_B_s_2^2 + 2*K_B_s_2*K_B_s_3 + 2*K_B_s_2*K_B_s_4 + 2*K_B_s_2*K_B_s_5 + 2*K_B_s_2 + K_B_s_3^2 + 2*K_B_s_3*K_B_s_4 + 2*K_B_s_3*K_B_s_5 + 2*K_B_s_3 + K_B_s_4^2 + 2*K_B_s_4*K_B_s_5 + 2*K_B_s_4 + K_B_s_5^2 + 2*K_B_s_5 + 1 and K_B_s_1 + K_B_s_2 + K_B_s_3 + K_B_s_4 + K_B_s_5 + (K_A^2*Ltot^2 - 2*K_A^2*Ltot*Rtot + K_A^2*Rtot^2 + 2*K_A*K_B_s_1*Ltot + 2*K_A*K_B_s_1*Rtot + 2*K_A*K_B_s_2*Ltot + 2*K_A*K_B_s_2*Rtot + 2*K_A*K_B_s_3*Ltot + 2*K_A*K_B_s_3*Rtot + 2*K_A*K_B_s_4*Ltot + 2*K_A*K_B_s_4*Rtot + 2*K_A*K_B_s_5*Ltot + 2*K_A*K_B_s_5*Rtot + 2*K_A*Ltot + 2*K_A*Rtot + K_B_s_1^2 + 2*K_B_s_1*K_B_s_2 + 2*K_B_s_1*K_B_s_3 + 2*K_B_s_1*K_B_s_4 + 2*K_B_s_1*K_B_s_5 + 2*K_B_s_1 + K_B_s_2^2 + 2*K_B_s_2*K_B_s_3 + 2*K_B_s_2*K_B_s_4 + 2*K_B_s_2*K_B_s_5 + 2*K_B_s_2 + K_B_s_3^2 + 2*K_B_s_3*K_B_s_4 + 2*K_B_s_3*K_B_s_5 + 2*K_B_s_3 + K_B_s_4^2 + 2*K_B_s_4*K_B_s_5 + 2*K_B_s_4 + K_B_s_5^2 + 2*K_B_s_5 + 1)^(1/2) + K_A*Rtot + 1 < K_A*Ltot, {-(K_B_s_1 + K_B_s_2 + K_B_s_3 + K_B_s_4 + K_B_s_5 + (K_A^2*Ltot^2 - 2*K_A^2*Ltot*Rtot + K_A^2*Rtot^2 + 2*K_A*K_B_s_1*Ltot + 2*K_A*K_B_s_1*Rtot + 2*K_A*K_B_s_2*Ltot + 2*K_A*K_B_s_2*Rtot + 2*K_A*K_B_s_3*Ltot + 2*K_A*K_B_s_3*Rtot + 2*K_A*K_B_s_4*Ltot + 2*K_A*K_B_s_4*Rtot + 2*K_A*K_B_s_5*Ltot + 2*K_A*K_B_s_5*Rtot + 2*K_A*Ltot + 2*K_A*Rtot + K_B_s_1^2 + 2*K_B_s_1*K_B_s_2 + 2*K_B_s_1*K_B_s_3 + 2*K_B_s_1*K_B_s_4 + 2*K_B_s_1*K_B_s_5 + 2*K_B_s_1 + K_B_s_2^2 + 2*K_B_s_2*K_B_s_3 + 2*K_B_s_2*K_B_s_4 + 2*K_B_s_2*K_B_s_5 + 2*K_B_s_2 + K_B_s_3^2 + 2*K_B_s_3*K_B_s_4 + 2*K_B_s_3*K_B_s_5 + 2*K_B_s_3 + K_B_s_4^2 + 2*K_B_s_4*K_B_s_5 + 2*K_B_s_4 + K_B_s_5^2 + 2*K_B_s_5 + 1)^(1/2) - K_A*Ltot + K_A*Rtot + 1)/(2*K_A)}], [K_B_s_1 + K_B_s_2 + K_B_s_3 + K_B_s_4 + K_B_s_5 - (K_A^2*Ltot^2 - 2*K_A^2*Ltot*Rtot + K_A^2*Rtot^2 + 2*K_A*K_B_s_1*Ltot + 2*K_A*K_B_s_1*Rtot + 2*K_A*K_B_s_2*Ltot + 2*K_A*K_B_s_2*Rtot + 2*K_A*K_B_s_3*Ltot + 2*K_A*K_B_s_3*Rtot + 2*K_A*K_B_s_4*Ltot + 2*K_A*K_B_s_4*Rtot + 2*K_A*K_B_s_5*Ltot + 2*K_A*K_B_s_5*Rtot + 2*K_A*Ltot + 2*K_A*Rtot + K_B_s_1^2 + 2*K_B_s_1*K_B_s_2 + 2*K_B_s_1*K_B_s_3 + 2*K_B_s_1*K_B_s_4 + 2*K_B_s_1*K_B_s_5 + 2*K_B_s_1 + K_B_s_2^2 + 2*K_B_s_2*K_B_s_3 + 2*K_B_s_2*K_B_s_4 + 2*K_B_s_2*K_B_s_5 + 2*K_B_s_2 + K_B_s_3^2 + 2*K_B_s_3*K_B_s_4 + 2*K_B_s_3*K_B_s_5 + 2*K_B_s_3 + K_B_s_4^2 + 2*K_B_s_4*K_B_s_5 + 2*K_B_s_4 + K_B_s_5^2 + 2*K_B_s_5 + 1)^(1/2) + K_A*Rtot + 1 < K_A*Ltot and (K_A^2*Ltot^2 - 2*K_A^2*Ltot*Rtot + K_A^2*Rtot^2 + 2*K_A*K_B_s_1*Ltot + 2*K_A*K_B_s_1*Rtot + 2*K_A*K_B_s_2*Ltot + 2*K_A*K_B_s_2*Rtot + 2*K_A*K_B_s_3*Ltot + 2*K_A*K_B_s_3*Rtot + 2*K_A*K_B_s_4*Ltot + 2*K_A*K_B_s_4*Rtot + 2*K_A*K_B_s_5*Ltot + 2*K_A*K_B_s_5*Rtot + 2*K_A*Ltot + 2*K_A*Rtot + K_B_s_1^2 + 2*K_B_s_1*K_B_s_2 + 2*K_B_s_1*K_B_s_3 + 2*K_B_s_1*K_B_s_4 + 2*K_B_s_1*K_B_s_5 + 2*K_B_s_1 + K_B_s_2^2 + 2*K_B_s_2*K_B_s_3 + 2*K_B_s_2*K_B_s_4 + 2*K_B_s_2*K_B_s_5 + 2*K_B_s_2 + K_B_s_3^2 + 2*K_B_s_3*K_B_s_4 + 2*K_B_s_3*K_B_s_5 + 2*K_B_s_3 + K_B_s_4^2 + 2*K_B_s_4*K_B_s_5 + 2*K_B_s_4 + K_B_s_5^2 + 2*K_B_s_5 + 1)^(1/2) + K_A*Ltot <> K_B_s_1 + K_B_s_2 + K_B_s_3 + K_B_s_4 + K_B_s_5 + K_A*Rtot + 1 and -(K_B_s_1 + K_B_s_2 + K_B_s_3 + K_B_s_4 + K_B_s_5 - (K_A^2*Ltot^2 - 2*K_A^2*Ltot*Rtot + K_A^2*Rtot^2 + 2*K_A*K_B_s_1*Ltot + 2*K_A*K_B_s_1*Rtot + 2*K_A*K_B_s_2*Ltot + 2*K_A*K_B_s_2*Rtot + 2*K_A*K_B_s_3*Ltot + 2*K_A*K_B_s_3*Rtot + 2*K_A*K_B_s_4*Ltot + 2*K_A*K_B_s_4*Rtot + 2*K_A*K_B_s_5*Ltot + 2*K_A*K_B_s_5*Rtot + 2*K_A*Ltot + 2*K_A*Rtot + K_B_s_1^2 + 2*K_B_s_1*K_B_s_2 + 2*K_B_s_1*K_B_s_3 + 2*K_B_s_1*K_B_s_4 + 2*K_B_s_1*K_B_s_5 + 2*K_B_s_1 + K_B_s_2^2 + 2*K_B_s_2*K_B_s_3 + 2*K_B_s_2*K_B_s_4 + 2*K_B_s_2*K_B_s_5 + 2*K_B_s_2 + K_B_s_3^2 + 2*K_B_s_3*K_B_s_4 + 2*K_B_s_3*K_B_s_5 + 2*K_B_s_3 + K_B_s_4^2 + 2*K_B_s_4*K_B_s_5 + 2*K_B_s_4 + K_B_s_5^2 + 2*K_B_s_5 + 1)^(1/2) - K_A*Ltot + K_A*Rtot + 1)/(2*K_A) < Ltot and (not -(K_B_s_1 + K_B_s_2 + K_B_s_3 + K_B_s_4 + K_B_s_5 + (K_A^2*Ltot^2 - 2*K_A^2*Ltot*Rtot + K_A^2*Rtot^2 + 2*K_A*K_B_s_1*Ltot + 2*K_A*K_B_s_1*Rtot + 2*K_A*K_B_s_2*Ltot + 2*K_A*K_B_s_2*Rtot + 2*K_A*K_B_s_3*Ltot + 2*K_A*K_B_s_3*Rtot + 2*K_A*K_B_s_4*Ltot + 2*K_A*K_B_s_4*Rtot + 2*K_A*K_B_s_5*Ltot + 2*K_A*K_B_s_5*Rtot + 2*K_A*Ltot + 2*K_A*Rtot + K_B_s_1^2 + 2*K_B_s_1*K_B_s_2 + 2*K_B_s_1*K_B_s_3 + 2*K_B_s_1*K_B_s_4 + 2*K_B_s_1*K_B_s_5 + 2*K_B_s_1 + K_B_s_2^2 + 2*K_B_s_2*K_B_s_3 + 2*K_B_s_2*K_B_s_4 + 2*K_B_s_2*K_B_s_5 + 2*K_B_s_2 + K_B_s_3^2 + 2*K_B_s_3*K_B_s_4 + 2*K_B_s_3*K_B_s_5 + 2*K_B_s_3 + K_B_s_4^2 + 2*K_B_s_4*K_B_s_5 + 2*K_B_s_4 + K_B_s_5^2 + 2*K_B_s_5 + 1)^(1/2) - K_A*Ltot + K_A*Rtot + 1)/(2*K_A) < Ltot and K_B_s_1 + K_B_s_2 + K_B_s_3 + K_B_s_4 + K_B_s_5 + (K_A^2*Ltot^2 - 2*K_A^2*Ltot*Rtot + K_A^2*Rtot^2 + 2*K_A*K_B_s_1*Ltot + 2*K_A*K_B_s_1*Rtot + 2*K_A*K_B_s_2*Ltot + 2*K_A*K_B_s_2*Rtot + 2*K_A*K_B_s_3*Ltot + 2*K_A*K_B_s_3*Rtot + 2*K_A*K_B_s_4*Ltot + 2*K_A*K_B_s_4*Rtot + 2*K_A*K_B_s_5*Ltot + 2*K_A*K_B_s_5*Rtot + 2*K_A*Ltot + 2*K_A*Rtot + K_B_s_1^2 + 2*K_B_s_1*K_B_s_2 + 2*K_B_s_1*K_B_s_3 + 2*K_B_s_1*K_B_s_4 + 2*K_B_s_1*K_B_s_5 + 2*K_B_s_1 + K_B_s_2^2 + 2*K_B_s_2*K_B_s_3 + 2*K_B_s_2*K_B_s_4 + 2*K_B_s_2*K_B_s_5 + 2*K_B_s_2 + K_B_s_3^2 + 2*K_B_s_3*K_B_s_4 + 2*K_B_s_3*K_B_s_5 + 2*K_B_s_3 + K_B_s_4^2 + 2*K_B_s_4*K_B_s_5 + 2*K_B_s_4 + K_B_s_5^2 + 2*K_B_s_5 + 1)^(1/2) + K_A*Rtot + 1 < K_A*Ltot or not K_B_s_1 + K_B_s_2 + K_B_s_3 + K_B_s_4 + K_B_s_5 + (K_A^2*Ltot^2 - 2*K_A^2*Ltot*Rtot + K_A^2*Rtot^2 + 2*K_A*K_B_s_1*Ltot + 2*K_A*K_B_s_1*Rtot + 2*K_A*K_B_s_2*Ltot + 2*K_A*K_B_s_2*Rtot + 2*K_A*K_B_s_3*Ltot + 2*K_A*K_B_s_3*Rtot + 2*K_A*K_B_s_4*Ltot + 2*K_A*K_B_s_4*Rtot + 2*K_A*K_B_s_5*Ltot + 2*K_A*K_B_s_5*Rtot + 2*K_A*Ltot + 2*K_A*Rtot + K_B_s_1^2 + 2*K_B_s_1*K_B_s_2 + 2*K_B_s_1*K_B_s_3 + 2*K_B_s_1*K_B_s_4 + 2*K_B_s_1*K_B_s_5 + 2*K_B_s_1 + K_B_s_2^2 + 2*K_B_s_2*K_B_s_3 + 2*K_B_s_2*K_B_s_4 + 2*K_B_s_2*K_B_s_5 + 2*K_B_s_2 + K_B_s_3^2 + 2*K_B_s_3*K_B_s_4 + 2*K_B_s_3*K_B_s_5 + 2*K_B_s_3 + K_B_s_4^2 + 2*K_B_s_4*K_B_s_5 + 2*K_B_s_4 + K_B_s_5^2 + 2*K_B_s_5 + 1)^(1/2) + K_A*Rtot + 1 <= K_A*Ltot) and 2*K_A^2*Ltot*Rtot <= K_A^2*Ltot^2 + K_A^2*Rtot^2 + 2*K_A*K_B_s_1*Ltot + 2*K_A*K_B_s_1*Rtot + 2*K_A*K_B_s_2*Ltot + 2*K_A*K_B_s_2*Rtot + 2*K_A*K_B_s_3*Ltot + 2*K_A*K_B_s_3*Rtot + 2*K_A*K_B_s_4*Ltot + 2*K_A*K_B_s_4*Rtot + 2*K_A*K_B_s_5*Ltot + 2*K_A*K_B_s_5*Rtot + 2*K_A*Ltot + 2*K_A*Rtot + K_B_s_1^2 + 2*K_B_s_1*K_B_s_2 + 2*K_B_s_1*K_B_s_3 + 2*K_B_s_1*K_B_s_4 + 2*K_B_s_1*K_B_s_5 + 2*K_B_s_1 + K_B_s_2^2 + 2*K_B_s_2*K_B_s_3 + 2*K_B_s_2*K_B_s_4 + 2*K_B_s_2*K_B_s_5 + 2*K_B_s_2 + K_B_s_3^2 + 2*K_B_s_3*K_B_s_4 + 2*K_B_s_3*K_B_s_5 + 2*K_B_s_3 + K_B_s_4^2 + 2*K_B_s_4*K_B_s_5 + 2*K_B_s_4 + K_B_s_5^2 + 2*K_B_s_5 + 1, {-(K_B_s_1 + K_B_s_2 + K_B_s_3 + K_B_s_4 + K_B_s_5 - (K_A^2*Ltot^2 - 2*K_A^2*Ltot*Rtot + K_A^2*Rtot^2 + 2*K_A*K_B_s_1*Ltot + 2*K_A*K_B_s_1*Rtot + 2*K_A*K_B_s_2*Ltot + 2*K_A*K_B_s_2*Rtot + 2*K_A*K_B_s_3*Ltot + 2*K_A*K_B_s_3*Rtot + 2*K_A*K_B_s_4*Ltot + 2*K_A*K_B_s_4*Rtot + 2*K_A*K_B_s_5*Ltot + 2*K_A*K_B_s_5*Rtot + 2*K_A*Ltot + 2*K_A*Rtot + K_B_s_1^2 + 2*K_B_s_1*K_B_s_2 + 2*K_B_s_1*K_B_s_3 + 2*K_B_s_1*K_B_s_4 + 2*K_B_s_1*K_B_s_5 + 2*K_B_s_1 + K_B_s_2^2 + 2*K_B_s_2*K_B_s_3 + 2*K_B_s_2*K_B_s_4 + 2*K_B_s_2*K_B_s_5 + 2*K_B_s_2 + K_B_s_3^2 + 2*K_B_s_3*K_B_s_4 + 2*K_B_s_3*K_B_s_5 + 2*K_B_s_3 + K_B_s_4^2 + 2*K_B_s_4*K_B_s_5 + 2*K_B_s_4 + K_B_s_5^2 + 2*K_B_s_5 + 1)^(1/2) - K_A*Ltot + K_A*Rtot + 1)/(2*K_A)}], [(K_B_s_1 + K_B_s_2 + K_B_s_3 + K_B_s_4 + K_B_s_5 - (K_A^2*Ltot^2 - 2*K_A^2*Ltot*Rtot + K_A^2*Rtot^2 + 2*K_A*K_B_s_1*Rtot + 2*K_A*K_B_s_2*Rtot + 2*K_A*K_B_s_3*Rtot + 2*K_A*K_B_s_4*Rtot + 2*K_A*K_B_s_5*Rtot + 2*K_A*Rtot + K_B_s_1^2 + 2*K_B_s_1*K_B_s_2 + 2*K_B_s_1*K_B_s_3 + 2*K_B_s_1*K_B_s_4 + 2*K_B_s_1*K_B_s_5 + 2*K_B_s_1 + K_B_s_2^2 + 2*K_B_s_2*K_B_s_3 + 2*K_B_s_2*K_B_s_4 + 2*K_B_s_2*K_B_s_5 + 2*K_B_s_2 + K_B_s_3^2 + 2*K_B_s_3*K_B_s_4 + 2*K_B_s_3*K_B_s_5 + 2*K_B_s_3 + K_B_s_4^2 + 2*K_B_s_4*K_B_s_5 + 2*K_B_s_4 + K_B_s_5^2 + 2*K_B_s_5 + 1)^(1/2) + K_A*Rtot + 1 < K_A*Ltot and (K_A^2*Ltot^2 - 2*K_A^2*Ltot*Rtot + K_A^2*Rtot^2 + 2*K_A*K_B_s_1*Ltot + 2*K_A*K_B_s_1*Rtot + 2*K_A*K_B_s_2*Ltot + 2*K_A*K_B_s_2*Rtot + 2*K_A*K_B_s_3*Ltot + 2*K_A*K_B_s_3*Rtot + 2*K_A*K_B_s_4*Ltot + 2*K_A*K_B_s_4*Rtot + 2*K_A*K_B_s_5*Ltot + 2*K_A*K_B_s_5*Rtot + 2*K_A*Ltot + 2*K_A*Rtot + K_B_s_1^2 + 2*K_B_s_1*K_B_s_2 + 2*K_B_s_1*K_B_s_3 + 2*K_B_s_1*K_B_s_4 + 2*K_B_s_1*K_B_s_5 + 2*K_B_s_1 + K_B_s_2^2 + 2*K_B_s_2*K_B_s_3 + 2*K_B_s_2*K_B_s_4 + 2*K_B_s_2*K_B_s_5 + 2*K_B_s_2 + K_B_s_3^2 + 2*K_B_s_3*K_B_s_4 + 2*K_B_s_3*K_B_s_5 + 2*K_B_s_3 + K_B_s_4^2 + 2*K_B_s_4*K_B_s_5 + 2*K_B_s_4 + K_B_s_5^2 + 2*K_B_s_5 + 1)^(1/2) + K_A*Ltot <> K_B_s_1 + K_B_s_2 + K_B_s_3 + K_B_s_4 + K_B_s_5 + K_A*Rtot + 1 and -(K_B_s_1 + K_B_s_2 + K_B_s_3 + K_B_s_4 + K_B_s_5 - (K_A^2*Ltot^2 - 2*K_A^2*Ltot*Rtot + K_A^2*Rtot^2 + 2*K_A*K_B_s_1*Rtot + 2*K_A*K_B_s_2*Rtot + 2*K_A*K_B_s_3*Rtot + 2*K_A*K_B_s_4*Rtot + 2*K_A*K_B_s_5*Rtot + 2*K_A*Rtot + K_B_s_1^2 + 2*K_B_s_1*K_B_s_2 + 2*K_B_s_1*K_B_s_3 + 2*K_B_s_1*K_B_s_4 + 2*K_B_s_1*K_B_s_5 + 2*K_B_s_1 + K_B_s_2^2 + 2*K_B_s_2*K_B_s_3 + 2*K_B_s_2*K_B_s_4 + 2*K_B_s_2*K_B_s_5 + 2*K_B_s_2 + K_B_s_3^2 + 2*K_B_s_3*K_B_s_4 + 2*K_B_s_3*K_B_s_5 + 2*K_B_s_3 + K_B_s_4^2 + 2*K_B_s_4*K_B_s_5 + 2*K_B_s_4 + K_B_s_5^2 + 2*K_B_s_5 + 1)^(1/2) - K_A*Ltot + K_A*Rtot + 1)/(2*K_A) < Ltot and K_B_s_1 + K_B_s_2 + K_B_s_3 + K_B_s_4 + K_B_s_5 + (K_A^2*Ltot^2 - 2*K_A^2*Ltot*Rtot + K_A^2*Rtot^2 + 2*K_A*K_B_s_1*Ltot + 2*K_A*K_B_s_1*Rtot + 2*K_A*K_B_s_2*Ltot + 2*K_A*K_B_s_2*Rtot + 2*K_A*K_B_s_3*Ltot + 2*K_A*K_B_s_3*Rtot + 2*K_A*K_B_s_4*Ltot + 2*K_A*K_B_s_4*Rtot + 2*K_A*K_B_s_5*Ltot + 2*K_A*K_B_s_5*Rtot + 2*K_A*Ltot + 2*K_A*Rtot + K_B_s_1^2 + 2*K_B_s_1*K_B_s_2 + 2*K_B_s_1*K_B_s_3 + 2*K_B_s_1*K_B_s_4 + 2*K_B_s_1*K_B_s_5 + 2*K_B_s_1 + K_B_s_2^2 + 2*K_B_s_2*K_B_s_3 + 2*K_B_s_2*K_B_s_4 + 2*K_B_s_2*K_B_s_5 + 2*K_B_s_2 + K_B_s_3^2 + 2*K_B_s_3*K_B_s_4 + 2*K_B_s_3*K_B_s_5 + 2*K_B_s_3 + K_B_s_4^2 + 2*K_B_s_4*K_B_s_5 + 2*K_B_s_4 + K_B_s_5^2 + 2*K_B_s_5 + 1)^(1/2) + K_A*Rtot + 1 = K_A*Ltot or -(K_B_s_1 + K_B_s_2 + K_B_s_3 + K_B_s_4 + K_B_s_5 + (K_A^2*Ltot^2 - 2*K_A^2*Ltot*Rtot + K_A^2*Rtot^2 + 2*K_A*K_B_s_1*Rtot + 2*K_A*K_B_s_2*Rtot + 2*K_A*K_B_s_3*Rtot + 2*K_A*K_B_s_4*Rtot + 2*K_A*K_B_s_5*Rtot + 2*K_A*Rtot + K_B_s_1^2 + 2*K_B_s_1*K_B_s_2 + 2*K_B_s_1*K_B_s_3 + 2*K_B_s_1*K_B_s_4 + 2*K_B_s_1*K_B_s_5 + 2*K_B_s_1 + K_B_s_2^2 + 2*K_B_s_2*K_B_s_3 + 2*K_B_s_2*K_B_s_4 + 2*K_B_s_2*K_B_s_5 + 2*K_B_s_2 + K_B_s_3^2 + 2*K_B_s_3*K_B_s_4 + 2*K_B_s_3*K_B_s_5 + 2*K_B_s_3 + K_B_s_4^2 + 2*K_B_s_4*K_B_s_5 + 2*K_B_s_4 + K_B_s_5^2 + 2*K_B_s_5 + 1)^(1/2) - K_A*Ltot + K_A*Rtot + 1)/(2*K_A) < Ltot and (K_A^2*Ltot^2 - 2*K_A^2*Ltot*Rtot + K_A^2*Rtot^2 + 2*K_A*K_B_s_1*Ltot + 2*K_A*K_B_s_1*Rtot + 2*K_A*K_B_s_2*Ltot + 2*K_A*K_B_s_2*Rtot + 2*K_A*K_B_s_3*Ltot + 2*K_A*K_B_s_3*Rtot + 2*K_A*K_B_s_4*Ltot + 2*K_A*K_B_s_4*Rtot + 2*K_A*K_B_s_5*Ltot + 2*K_A*K_B_s_5*Rtot + 2*K_A*Ltot + 2*K_A*Rtot + K_B_s_1^2 + 2*K_B_s_1*K_B_s_2 + 2*K_B_s_1*K_B_s_3 + 2*K_B_s_1*K_B_s_4 + 2*K_B_s_1*K_B_s_5 + 2*K_B_s_1 + K_B_s_2^2 + 2*K_B_s_2*K_B_s_3 + 2*K_B_s_2*K_B_s_4 + 2*K_B_s_2*K_B_s_5 + 2*K_B_s_2 + K_B_s_3^2 + 2*K_B_s_3*K_B_s_4 + 2*K_B_s_3*K_B_s_5 + 2*K_B_s_3 + K_B_s_4^2 + 2*K_B_s_4*K_B_s_5 + 2*K_B_s_4 + K_B_s_5^2 + 2*K_B_s_5 + 1)^(1/2) + K_A*Ltot = K_B_s_1 + K_B_s_2 + K_B_s_3 + K_B_s_4 + K_B_s_5 + K_A*Rtot + 1 and K_B_s_1 + K_B_s_2 + K_B_s_3 + K_B_s_4 + K_B_s_5 + (K_A^2*Ltot^2 - 2*K_A^2*Ltot*Rtot + K_A^2*Rtot^2 + 2*K_A*K_B_s_1*Rtot + 2*K_A*K_B_s_2*Rtot + 2*K_A*K_B_s_3*Rtot + 2*K_A*K_B_s_4*Rtot + 2*K_A*K_B_s_5*Rtot + 2*K_A*Rtot + K_B_s_1^2 + 2*K_B_s_1*K_B_s_2 + 2*K_B_s_1*K_B_s_3 + 2*K_B_s_1*K_B_s_4 + 2*K_B_s_1*K_B_s_5 + 2*K_B_s_1 + K_B_s_2^2 + 2*K_B_s_2*K_B_s_3 + 2*K_B_s_2*K_B_s_4 + 2*K_B_s_2*K_B_s_5 + 2*K_B_s_2 + K_B_s_3^2 + 2*K_B_s_3*K_B_s_4 + 2*K_B_s_3*K_B_s_5 + 2*K_B_s_3 + K_B_s_4^2 + 2*K_B_s_4*K_B_s_5 + 2*K_B_s_4 + K_B_s_5^2 + 2*K_B_s_5 + 1)^(1/2) + K_A*Rtot + 1 < K_A*Ltot and K_B_s_1 + K_B_s_2 + K_B_s_3 + K_B_s_4 + K_B_s_5 + (K_A^2*Ltot^2 - 2*K_A^2*Ltot*Rtot + K_A^2*Rtot^2 + 2*K_A*K_B_s_1*Ltot + 2*K_A*K_B_s_1*Rtot + 2*K_A*K_B_s_2*Ltot + 2*K_A*K_B_s_2*Rtot + 2*K_A*K_B_s_3*Ltot + 2*K_A*K_B_s_3*Rtot + 2*K_A*K_B_s_4*Ltot + 2*K_A*K_B_s_4*Rtot + 2*K_A*K_B_s_5*Ltot + 2*K_A*K_B_s_5*Rtot + 2*K_A*Ltot + 2*K_A*Rtot + K_B_s_1^2 + 2*K_B_s_1*K_B_s_2 + 2*K_B_s_1*K_B_s_3 + 2*K_B_s_1*K_B_s_4 + 2*K_B_s_1*K_B_s_5 + 2*K_B_s_1 + K_B_s_2^2 + 2*K_B_s_2*K_B_s_3 + 2*K_B_s_2*K_B_s_4 + 2*K_B_s_2*K_B_s_5 + 2*K_B_s_2 + K_B_s_3^2 + 2*K_B_s_3*K_B_s_4 + 2*K_B_s_3*K_B_s_5 + 2*K_B_s_3 + K_B_s_4^2 + 2*K_B_s_4*K_B_s_5 + 2*K_B_s_4 + K_B_s_5^2 + 2*K_B_s_5 + 1)^(1/2) + K_A*Rtot + 1 <> K_A*Ltot) and K_A^2*Ltot^2 + K_A^2*Rtot^2 + 2*K_A*K_B_s_1*Rtot + 2*K_A*K_B_s_2*Rtot + 2*K_A*K_B_s_3*Rtot + 2*K_A*K_B_s_4*Rtot + 2*K_A*K_B_s_5*Rtot + 2*K_A*Rtot + K_B_s_1^2 + 2*K_B_s_1*K_B_s_2 + 2*K_B_s_1*K_B_s_3 + 2*K_B_s_1*K_B_s_4 + 2*K_B_s_1*K_B_s_5 + 2*K_B_s_1 + K_B_s_2^2 + 2*K_B_s_2*K_B_s_3 + 2*K_B_s_2*K_B_s_4 + 2*K_B_s_2*K_B_s_5 + 2*K_B_s_2 + K_B_s_3^2 + 2*K_B_s_3*K_B_s_4 + 2*K_B_s_3*K_B_s_5 + 2*K_B_s_3 + K_B_s_4^2 + 2*K_B_s_4*K_B_s_5 + 2*K_B_s_4 + K_B_s_5^2 + 2*K_B_s_5 + 1 < 2*K_A^2*Ltot*Rtot or (K_A^2*Ltot^2 - 2*K_A^2*Ltot*Rtot + K_A^2*Rtot^2 + 2*K_A*K_B_s_1*Ltot + 2*K_A*K_B_s_1*Rtot + 2*K_A*K_B_s_2*Ltot + 2*K_A*K_B_s_2*Rtot + 2*K_A*K_B_s_3*Ltot + 2*K_A*K_B_s_3*Rtot + 2*K_A*K_B_s_4*Ltot + 2*K_A*K_B_s_4*Rtot + 2*K_A*K_B_s_5*Ltot + 2*K_A*K_B_s_5*Rtot + 2*K_A*Ltot + 2*K_A*Rtot + K_B_s_1^2 + 2*K_B_s_1*K_B_s_2 + 2*K_B_s_1*K_B_s_3 + 2*K_B_s_1*K_B_s_4 + 2*K_B_s_1*K_B_s_5 + 2*K_B_s_1 + K_B_s_2^2 + 2*K_B_s_2*K_B_s_3 + 2*K_B_s_2*K_B_s_4 + 2*K_B_s_2*K_B_s_5 + 2*K_B_s_2 + K_B_s_3^2 + 2*K_B_s_3*K_B_s_4 + 2*K_B_s_3*K_B_s_5 + 2*K_B_s_3 + K_B_s_4^2 + 2*K_B_s_4*K_B_s_5 + 2*K_B_s_4 + K_B_s_5^2 + 2*K_B_s_5 + 1)^(1/2) + K_A*Ltot = K_B_s_1 + K_B_s_2 + K_B_s_3 + K_B_s_4 + K_B_s_5 + K_A*Rtot + 1 and (not K_B_s_1 + K_B_s_2 + K_B_s_3 + K_B_s_4 + K_B_s_5 + (K_A^2*Ltot^2 - 2*K_A^2*Ltot*Rtot + K_A^2*Rtot^2 + 2*K_A*K_B_s_1*Rtot + 2*K_A*K_B_s_2*Rtot + 2*K_A*K_B_s_3*Rtot + 2*K_A*K_B_s_4*Rtot + 2*K_A*K_B_s_5*Rtot + 2*K_A*Rtot + K_B_s_1^2 + 2*K_B_s_1*K_B_s_2 + 2*K_B_s_1*K_B_s_3 + 2*K_B_s_1*K_B_s_4 + 2*K_B_s_1*K_B_s_5 + 2*K_B_s_1 + K_B_s_2^2 + 2*K_B_s_2*K_B_s_3 + 2*K_B_s_2*K_B_s_4 + 2*K_B_s_2*K_B_s_5 + 2*K_B_s_2 + K_B_s_3^2 + 2*K_B_s_3*K_B_s_4 + 2*K_B_s_3*K_B_s_5 + 2*K_B_s_3 + K_B_s_4^2 + 2*K_B_s_4*K_B_s_5 + 2*K_B_s_4 + K_B_s_5^2 + 2*K_B_s_5 + 1)^(1/2) + K_A*Rtot + 1 < K_A*Ltot or K_B_s_1 + K_B_s_2 + K_B_s_3 + K_B_s_4 + K_B_s_5 + (K_A^2*Ltot^2 - 2*K_A^2*Ltot*Rtot + K_A^2*Rtot^2 + 2*K_A*K_B_s_1*Ltot + 2*K_A*K_B_s_1*Rtot + 2*K_A*K_B_s_2*Ltot + 2*K_A*K_B_s_2*Rtot + 2*K_A*K_B_s_3*Ltot + 2*K_A*K_B_s_3*Rtot + 2*K_A*K_B_s_4*Ltot + 2*K_A*K_B_s_4*Rtot + 2*K_A*K_B_s_5*Ltot + 2*K_A*K_B_s_5*Rtot + 2*K_A*Ltot + 2*K_A*Rtot + K_B_s_1^2 + 2*K_B_s_1*K_B_s_2 + 2*K_B_s_1*K_B_s_3 + 2*K_B_s_1*K_B_s_4 + 2*K_B_s_1*K_B_s_5 + 2*K_B_s_1 + K_B_s_2^2 + 2*K_B_s_2*K_B_s_3 + 2*K_B_s_2*K_B_s_4 + 2*K_B_s_2*K_B_s_5 + 2*K_B_s_2 + K_B_s_3^2 + 2*K_B_s_3*K_B_s_4 + 2*K_B_s_3*K_B_s_5 + 2*K_B_s_3 + K_B_s_4^2 + 2*K_B_s_4*K_B_s_5 + 2*K_B_s_4 + K_B_s_5^2 + 2*K_B_s_5 + 1)^(1/2) + K_A*Rtot + 1 = K_A*Ltot) or not K_B_s_1 + K_B_s_2 + K_B_s_3 + K_B_s_4 + K_B_s_5 - (K_A^2*Ltot^2 - 2*K_A^2*Ltot*Rtot + K_A^2*Rtot^2 + 2*K_A*K_B_s_1*Rtot + 2*K_A*K_B_s_2*Rtot + 2*K_A*K_B_s_3*Rtot + 2*K_A*K_B_s_4*Rtot + 2*K_A*K_B_s_5*Rtot + 2*K_A*Rtot + K_B_s_1^2 + 2*K_B_s_1*K_B_s_2 + 2*K_B_s_1*K_B_s_3 + 2*K_B_s_1*K_B_s_4 + 2*K_B_s_1*K_B_s_5 + 2*K_B_s_1 + K_B_s_2^2 + 2*K_B_s_2*K_B_s_3 + 2*K_B_s_2*K_B_s_4 + 2*K_B_s_2*K_B_s_5 + 2*K_B_s_2 + K_B_s_3^2 + 2*K_B_s_3*K_B_s_4 + 2*K_B_s_3*K_B_s_5 + 2*K_B_s_3 + K_B_s_4^2 + 2*K_B_s_4*K_B_s_5 + 2*K_B_s_4 + K_B_s_5^2 + 2*K_B_s_5 + 1)^(1/2) + K_A*Rtot + 1 < K_A*Ltot and (K_A^2*Ltot^2 - 2*K_A^2*Ltot*Rtot + K_A^2*Rtot^2 + 2*K_A*K_B_s_1*Ltot + 2*K_A*K_B_s_1*Rtot + 2*K_A*K_B_s_2*Ltot + 2*K_A*K_B_s_2*Rtot + 2*K_A*K_B_s_3*Ltot + 2*K_A*K_B_s_3*Rtot + 2*K_A*K_B_s_4*Ltot + 2*K_A*K_B_s_4*Rtot + 2*K_A*K_B_s_5*Ltot + 2*K_A*K_B_s_5*Rtot + 2*K_A*Ltot + 2*K_A*Rtot + K_B_s_1^2 + 2*K_B_s_1*K_B_s_2 + 2*K_B_s_1*K_B_s_3 + 2*K_B_s_1*K_B_s_4 + 2*K_B_s_1*K_B_s_5 + 2*K_B_s_1 + K_B_s_2^2 + 2*K_B_s_2*K_B_s_3 + 2*K_B_s_2*K_B_s_4 + 2*K_B_s_2*K_B_s_5 + 2*K_B_s_2 + K_B_s_3^2 + 2*K_B_s_3*K_B_s_4 + 2*K_B_s_3*K_B_s_5 + 2*K_B_s_3 + K_B_s_4^2 + 2*K_B_s_4*K_B_s_5 + 2*K_B_s_4 + K_B_s_5^2 + 2*K_B_s_5 + 1)^(1/2) + K_A*Ltot <> K_B_s_1 + K_B_s_2 + K_B_s_3 + K_B_s_4 + K_B_s_5 + K_A*Rtot + 1 and K_B_s_1 + K_B_s_2 + K_B_s_3 + K_B_s_4 + K_B_s_5 + (K_A^2*Ltot^2 - 2*K_A^2*Ltot*Rtot + K_A^2*Rtot^2 + 2*K_A*K_B_s_1*Ltot + 2*K_A*K_B_s_1*Rtot + 2*K_A*K_B_s_2*Ltot + 2*K_A*K_B_s_2*Rtot + 2*K_A*K_B_s_3*Ltot + 2*K_A*K_B_s_3*Rtot + 2*K_A*K_B_s_4*Ltot + 2*K_A*K_B_s_4*Rtot + 2*K_A*K_B_s_5*Ltot + 2*K_A*K_B_s_5*Rtot + 2*K_A*Ltot + 2*K_A*Rtot + K_B_s_1^2 + 2*K_B_s_1*K_B_s_2 + 2*K_B_s_1*K_B_s_3 + 2*K_B_s_1*K_B_s_4 + 2*K_B_s_1*K_B_s_5 + 2*K_B_s_1 + K_B_s_2^2 + 2*K_B_s_2*K_B_s_3 + 2*K_B_s_2*K_B_s_4 + 2*K_B_s_2*K_B_s_5 + 2*K_B_s_2 + K_B_s_3^2 + 2*K_B_s_3*K_B_s_4 + 2*K_B_s_3*K_B_s_5 + 2*K_B_s_3 + K_B_s_4^2 + 2*K_B_s_4*K_B_s_5 + 2*K_B_s_4 + K_B_s_5^2 + 2*K_B_s_5 + 1)^(1/2) + K_A*Rtot + 1 = K_A*Ltot or not K_B_s_1 + K_B_s_2 + K_B_s_3 + K_B_s_4 + K_B_s_5 - (K_A^2*Ltot^2 - 2*K_A^2*Ltot*Rtot + K_A^2*Rtot^2 + 2*K_A*K_B_s_1*Ltot + 2*K_A*K_B_s_1*Rtot + 2*K_A*K_B_s_2*Ltot + 2*K_A*K_B_s_2*Rtot + 2*K_A*K_B_s_3*Ltot + 2*K_A*K_B_s_3*Rtot + 2*K_A*K_B_s_4*Ltot + 2*K_A*K_B_s_4*Rtot + 2*K_A*K_B_s_5*Ltot + 2*K_A*K_B_s_5*Rtot + 2*K_A*Ltot + 2*K_A*Rtot + K_B_s_1^2 + 2*K_B_s_1*K_B_s_2 + 2*K_B_s_1*K_B_s_3 + 2*K_B_s_1*K_B_s_4 + 2*K_B_s_1*K_B_s_5 + 2*K_B_s_1 + K_B_s_2^2 + 2*K_B_s_2*K_B_s_3 + 2*K_B_s_2*K_B_s_4 + 2*K_B_s_2*K_B_s_5 + 2*K_B_s_2 + K_B_s_3^2 + 2*K_B_s_3*K_B_s_4 + 2*K_B_s_3*K_B_s_5 + 2*K_B_s_3 + K_B_s_4^2 + 2*K_B_s_4*K_B_s_5 + 2*K_B_s_4 + K_B_s_5^2 + 2*K_B_s_5 + 1)^(1/2) + K_A*Rtot + 1 < K_A*Ltot and (K_A^2*Ltot^2 - 2*K_A^2*Ltot*Rtot + K_A^2*Rtot^2 + 2*K_A*K_B_s_1*Ltot + 2*K_A*K_B_s_1*Rtot + 2*K_A*K_B_s_2*Ltot + 2*K_A*K_B_s_2*Rtot + 2*K_A*K_B_s_3*Ltot + 2*K_A*K_B_s_3*Rtot + 2*K_A*K_B_s_4*Ltot + 2*K_A*K_B_s_4*Rtot + 2*K_A*K_B_s_5*Ltot + 2*K_A*K_B_s_5*Rtot + 2*K_A*Ltot + 2*K_A*Rtot + K_B_s_1^2 + 2*K_B_s_1*K_B_s_2 + 2*K_B_s_1*K_B_s_3 + 2*K_B_s_1*K_B_s_4 + 2*K_B_s_1*K_B_s_5 + 2*K_B_s_1 + K_B_s_2^2 + 2*K_B_s_2*K_B_s_3 + 2*K_B_s_2*K_B_s_4 + 2*K_B_s_2*K_B_s_5 + 2*K_B_s_2 + K_B_s_3^2 + 2*K_B_s_3*K_B_s_4 + 2*K_B_s_3*K_B_s_5 + 2*K_B_s_3 + K_B_s_4^2 + 2*K_B_s_4*K_B_s_5 + 2*K_B_s_4 + K_B_s_5^2 + 2*K_B_s_5 + 1)^(1/2) + K_A*Ltot <> K_B_s_1 + K_B_s_2 + K_B_s_3 + K_B_s_4 + K_B_s_5 + K_A*Rtot + 1 and not K_B_s_1 + K_B_s_2 + K_B_s_3 + K_B_s_4 + K_B_s_5 + (K_A^2*Ltot^2 - 2*K_A^2*Ltot*Rtot + K_A^2*Rtot^2 + 2*K_A*K_B_s_1*Ltot + 2*K_A*K_B_s_1*Rtot + 2*K_A*K_B_s_2*Ltot + 2*K_A*K_B_s_2*Rtot + 2*K_A*K_B_s_3*Ltot + 2*K_A*K_B_s_3*Rtot + 2*K_A*K_B_s_4*Ltot + 2*K_A*K_B_s_4*Rtot + 2*K_A*K_B_s_5*Ltot + 2*K_A*K_B_s_5*Rtot + 2*K_A*Ltot + 2*K_A*Rtot + K_B_s_1^2 + 2*K_B_s_1*K_B_s_2 + 2*K_B_s_1*K_B_s_3 + 2*K_B_s_1*K_B_s_4 + 2*K_B_s_1*K_B_s_5 + 2*K_B_s_1 + K_B_s_2^2 + 2*K_B_s_2*K_B_s_3 + 2*K_B_s_2*K_B_s_4 + 2*K_B_s_2*K_B_s_5 + 2*K_B_s_2 + K_B_s_3^2 + 2*K_B_s_3*K_B_s_4 + 2*K_B_s_3*K_B_s_5 + 2*K_B_s_3 + K_B_s_4^2 + 2*K_B_s_4*K_B_s_5 + 2*K_B_s_4 + K_B_s_5^2 + 2*K_B_s_5 + 1)^(1/2) + K_A*Rtot + 1 <= K_A*Ltot or not -(K_B_s_1 + K_B_s_2 + K_B_s_3 + K_B_s_4 + K_B_s_5 + (K_A^2*Ltot^2 - 2*K_A^2*Ltot*Rtot + K_A^2*Rtot^2 + 2*K_A*K_B_s_1*Ltot + 2*K_A*K_B_s_1*Rtot + 2*K_A*K_B_s_2*Ltot + 2*K_A*K_B_s_2*Rtot + 2*K_A*K_B_s_3*Ltot + 2*K_A*K_B_s_3*Rtot + 2*K_A*K_B_s_4*Ltot + 2*K_A*K_B_s_4*Rtot + 2*K_A*K_B_s_5*Ltot + 2*K_A*K_B_s_5*Rtot + 2*K_A*Ltot + 2*K_A*Rtot + K_B_s_1^2 + 2*K_B_s_1*K_B_s_2 + 2*K_B_s_1*K_B_s_3 + 2*K_B_s_1*K_B_s_4 + 2*K_B_s_1*K_B_s_5 + 2*K_B_s_1 + K_B_s_2^2 + 2*K_B_s_2*K_B_s_3 + 2*K_B_s_2*K_B_s_4 + 2*K_B_s_2*K_B_s_5 + 2*K_B_s_2 + K_B_s_3^2 + 2*K_B_s_3*K_B_s_4 + 2*K_B_s_3*K_B_s_5 + 2*K_B_s_3 + K_B_s_4^2 + 2*K_B_s_4*K_B_s_5 + 2*K_B_s_4 + K_B_s_5^2 + 2*K_B_s_5 + 1)^(1/2) - K_A*Ltot + K_A*Rtot + 1)/(2*K_A) < Ltot and not K_B_s_1 + K_B_s_2 + K_B_s_3 + K_B_s_4 + K_B_s_5 - (K_A^2*Ltot^2 - 2*K_A^2*Ltot*Rtot + K_A^2*Rtot^2 + 2*K_A*K_B_s_1*Ltot + 2*K_A*K_B_s_1*Rtot + 2*K_A*K_B_s_2*Ltot + 2*K_A*K_B_s_2*Rtot + 2*K_A*K_B_s_3*Ltot + 2*K_A*K_B_s_3*Rtot + 2*K_A*K_B_s_4*Ltot + 2*K_A*K_B_s_4*Rtot + 2*K_A*K_B_s_5*Ltot + 2*K_A*K_B_s_5*Rtot + 2*K_A*Ltot + 2*K_A*Rtot + K_B_s_1^2 + 2*K_B_s_1*K_B_s_2 + 2*K_B_s_1*K_B_s_3 + 2*K_B_s_1*K_B_s_4 + 2*K_B_s_1*K_B_s_5 + 2*K_B_s_1 + K_B_s_2^2 + 2*K_B_s_2*K_B_s_3 + 2*K_B_s_2*K_B_s_4 + 2*K_B_s_2*K_B_s_5 + 2*K_B_s_2 + K_B_s_3^2 + 2*K_B_s_3*K_B_s_4 + 2*K_B_s_3*K_B_s_5 + 2*K_B_s_3 + K_B_s_4^2 + 2*K_B_s_4*K_B_s_5 + 2*K_B_s_4 + K_B_s_5^2 + 2*K_B_s_5 + 1)^(1/2) + K_A*Rtot + 1 < K_A*Ltot and (K_A^2*Ltot^2 - 2*K_A^2*Ltot*Rtot + K_A^2*Rtot^2 + 2*K_A*K_B_s_1*Ltot + 2*K_A*K_B_s_1*Rtot + 2*K_A*K_B_s_2*Ltot + 2*K_A*K_B_s_2*Rtot + 2*K_A*K_B_s_3*Ltot + 2*K_A*K_B_s_3*Rtot + 2*K_A*K_B_s_4*Ltot + 2*K_A*K_B_s_4*Rtot + 2*K_A*K_B_s_5*Ltot + 2*K_A*K_B_s_5*Rtot + 2*K_A*Ltot + 2*K_A*Rtot + K_B_s_1^2 + 2*K_B_s_1*K_B_s_2 + 2*K_B_s_1*K_B_s_3 + 2*K_B_s_1*K_B_s_4 + 2*K_B_s_1*K_B_s_5 + 2*K_B_s_1 + K_B_s_2^2 + 2*K_B_s_2*K_B_s_3 + 2*K_B_s_2*K_B_s_4 + 2*K_B_s_2*K_B_s_5 + 2*K_B_s_2 + K_B_s_3^2 + 2*K_B_s_3*K_B_s_4 + 2*K_B_s_3*K_B_s_5 + 2*K_B_s_3 + K_B_s_4^2 + 2*K_B_s_4*K_B_s_5 + 2*K_B_s_4 + K_B_s_5^2 + 2*K_B_s_5 + 1)^(1/2) + K_A*Ltot <> K_B_s_1 + K_B_s_2 + K_B_s_3 + K_B_s_4 + K_B_s_5 + K_A*Rtot + 1 and K_B_s_1 + K_B_s_2 + K_B_s_3 + K_B_s_4 + K_B_s_5 + (K_A^2*Ltot^2 - 2*K_A^2*Ltot*Rtot + K_A^2*Rtot^2 + 2*K_A*K_B_s_1*Ltot + 2*K_A*K_B_s_1*Rtot + 2*K_A*K_B_s_2*Ltot + 2*K_A*K_B_s_2*Rtot + 2*K_A*K_B_s_3*Ltot + 2*K_A*K_B_s_3*Rtot + 2*K_A*K_B_s_4*Ltot + 2*K_A*K_B_s_4*Rtot + 2*K_A*K_B_s_5*Ltot + 2*K_A*K_B_s_5*Rtot + 2*K_A*Ltot + 2*K_A*Rtot + K_B_s_1^2 + 2*K_B_s_1*K_B_s_2 + 2*K_B_s_1*K_B_s_3 + 2*K_B_s_1*K_B_s_4 + 2*K_B_s_1*K_B_s_5 + 2*K_B_s_1 + K_B_s_2^2 + 2*K_B_s_2*K_B_s_3 + 2*K_B_s_2*K_B_s_4 + 2*K_B_s_2*K_B_s_5 + 2*K_B_s_2 + K_B_s_3^2 + 2*K_B_s_3*K_B_s_4 + 2*K_B_s_3*K_B_s_5 + 2*K_B_s_3 + K_B_s_4^2 + 2*K_B_s_4*K_B_s_5 + 2*K_B_s_4 + K_B_s_5^2 + 2*K_B_s_5 + 1)^(1/2) + K_A*Rtot + 1 < K_A*Ltot or K_B_s_1 + K_B_s_2 + K_B_s_3 + K_B_s_4 + K_B_s_5 - (K_A^2*Ltot^2 - 2*K_A^2*Ltot*Rtot + K_A^2*Rtot^2 + 2*K_A*K_B_s_1*Ltot + 2*K_A*K_B_s_1*Rtot + 2*K_A*K_B_s_2*Ltot + 2*K_A*K_B_s_2*Rtot + 2*K_A*K_B_s_3*Ltot + 2*K_A*K_B_s_3*Rtot + 2*K_A*K_B_s_4*Ltot + 2*K_A*K_B_s_4*Rtot + 2*K_A*K_B_s_5*Ltot + 2*K_A*K_B_s_5*Rtot + 2*K_A*Ltot + 2*K_A*Rtot + K_B_s_1^2 + 2*K_B_s_1*K_B_s_2 + 2*K_B_s_1*K_B_s_3 + 2*K_B_s_1*K_B_s_4 + 2*K_B_s_1*K_B_s_5 + 2*K_B_s_1 + K_B_s_2^2 + 2*K_B_s_2*K_B_s_3 + 2*K_B_s_2*K_B_s_4 + 2*K_B_s_2*K_B_s_5 + 2*K_B_s_2 + K_B_s_3^2 + 2*K_B_s_3*K_B_s_4 + 2*K_B_s_3*K_B_s_5 + 2*K_B_s_3 + K_B_s_4^2 + 2*K_B_s_4*K_B_s_5 + 2*K_B_s_4 + K_B_s_5^2 + 2*K_B_s_5 + 1)^(1/2) + K_A*Rtot + 1 < K_A*Ltot and (K_A^2*Ltot^2 - 2*K_A^2*Ltot*Rtot + K_A^2*Rtot^2 + 2*K_A*K_B_s_1*Ltot + 2*K_A*K_B_s_1*Rtot + 2*K_A*K_B_s_2*Ltot + 2*K_A*K_B_s_2*Rtot + 2*K_A*K_B_s_3*Ltot + 2*K_A*K_B_s_3*Rtot + 2*K_A*K_B_s_4*Ltot + 2*K_A*K_B_s_4*Rtot + 2*K_A*K_B_s_5*Ltot + 2*K_A*K_B_s_5*Rtot + 2*K_A*Ltot + 2*K_A*Rtot + K_B_s_1^2 + 2*K_B_s_1*K_B_s_2 + 2*K_B_s_1*K_B_s_3 + 2*K_B_s_1*K_B_s_4 + 2*K_B_s_1*K_B_s_5 + 2*K_B_s_1 + K_B_s_2^2 + 2*K_B_s_2*K_B_s_3 + 2*K_B_s_2*K_B_s_4 + 2*K_B_s_2*K_B_s_5 + 2*K_B_s_2 + K_B_s_3^2 + 2*K_B_s_3*K_B_s_4 + 2*K_B_s_3*K_B_s_5 + 2*K_B_s_3 + K_B_s_4^2 + 2*K_B_s_4*K_B_s_5 + 2*K_B_s_4 + K_B_s_5^2 + 2*K_B_s_5 + 1)^(1/2) + K_A*Ltot <> K_B_s_1 + K_B_s_2 + K_B_s_3 + K_B_s_4 + K_B_s_5 + K_A*Rtot + 1 and not -(K_B_s_1 + K_B_s_2 + K_B_s_3 + K_B_s_4 + K_B_s_5 - (K_A^2*Ltot^2 - 2*K_A^2*Ltot*Rtot + K_A^2*Rtot^2 + 2*K_A*K_B_s_1*Ltot + 2*K_A*K_B_s_1*Rtot + 2*K_A*K_B_s_2*Ltot + 2*K_A*K_B_s_2*Rtot + 2*K_A*K_B_s_3*Ltot + 2*K_A*K_B_s_3*Rtot + 2*K_A*K_B_s_4*Ltot + 2*K_A*K_B_s_4*Rtot + 2*K_A*K_B_s_5*Ltot + 2*K_A*K_B_s_5*Rtot + 2*K_A*Ltot + 2*K_A*Rtot + K_B_s_1^2 + 2*K_B_s_1*K_B_s_2 + 2*K_B_s_1*K_B_s_3 + 2*K_B_s_1*K_B_s_4 + 2*K_B_s_1*K_B_s_5 + 2*K_B_s_1 + K_B_s_2^2 + 2*K_B_s_2*K_B_s_3 + 2*K_B_s_2*K_B_s_4 + 2*K_B_s_2*K_B_s_5 + 2*K_B_s_2 + K_B_s_3^2 + 2*K_B_s_3*K_B_s_4 + 2*K_B_s_3*K_B_s_5 + 2*K_B_s_3 + K_B_s_4^2 + 2*K_B_s_4*K_B_s_5 + 2*K_B_s_4 + K_B_s_5^2 + 2*K_B_s_5 + 1)^(1/2) - K_A*Ltot + K_A*Rtot + 1)/(2*K_A) < Ltot and not K_B_s_1 + K_B_s_2 + K_B_s_3 + K_B_s_4 + K_B_s_5 + (K_A^2*Ltot^2 - 2*K_A^2*Ltot*Rtot + K_A^2*Rtot^2 + 2*K_A*K_B_s_1*Ltot + 2*K_A*K_B_s_1*Rtot + 2*K_A*K_B_s_2*Ltot + 2*K_A*K_B_s_2*Rtot + 2*K_A*K_B_s_3*Ltot + 2*K_A*K_B_s_3*Rtot + 2*K_A*K_B_s_4*Ltot + 2*K_A*K_B_s_4*Rtot + 2*K_A*K_B_s_5*Ltot + 2*K_A*K_B_s_5*Rtot + 2*K_A*Ltot + 2*K_A*Rtot + K_B_s_1^2 + 2*K_B_s_1*K_B_s_2 + 2*K_B_s_1*K_B_s_3 + 2*K_B_s_1*K_B_s_4 + 2*K_B_s_1*K_B_s_5 + 2*K_B_s_1 + K_B_s_2^2 + 2*K_B_s_2*K_B_s_3 + 2*K_B_s_2*K_B_s_4 + 2*K_B_s_2*K_B_s_5 + 2*K_B_s_2 + K_B_s_3^2 + 2*K_B_s_3*K_B_s_4 + 2*K_B_s_3*K_B_s_5 + 2*K_B_s_3 + K_B_s_4^2 + 2*K_B_s_4*K_B_s_5 + 2*K_B_s_4 + K_B_s_5^2 + 2*K_B_s_5 + 1)^(1/2) + K_A*Rtot + 1 <= K_A*Ltot or K_B_s_1 + K_B_s_2 + K_B_s_3 + K_B_s_4 + K_B_s_5 - (K_A^2*Ltot^2 - 2*K_A^2*Ltot*Rtot + K_A^2*Rtot^2 + 2*K_A*K_B_s_1*Rtot + 2*K_A*K_B_s_2*Rtot + 2*K_A*K_B_s_3*Rtot + 2*K_A*K_B_s_4*Rtot + 2*K_A*K_B_s_5*Rtot + 2*K_A*Rtot + K_B_s_1^2 + 2*K_B_s_1*K_B_s_2 + 2*K_B_s_1*K_B_s_3 + 2*K_B_s_1*K_B_s_4 + 2*K_B_s_1*K_B_s_5 + 2*K_B_s_1 + K_B_s_2^2 + 2*K_B_s_2*K_B_s_3 + 2*K_B_s_2*K_B_s_4 + 2*K_B_s_2*K_B_s_5 + 2*K_B_s_2 + K_B_s_3^2 + 2*K_B_s_3*K_B_s_4 + 2*K_B_s_3*K_B_s_5 + 2*K_B_s_3 + K_B_s_4^2 + 2*K_B_s_4*K_B_s_5 + 2*K_B_s_4 + K_B_s_5^2 + 2*K_B_s_5 + 1)^(1/2) + K_A*Rtot + 1 < K_A*Ltot and (K_A^2*Ltot^2 - 2*K_A^2*Ltot*Rtot + K_A^2*Rtot^2 + 2*K_A*K_B_s_1*Ltot + 2*K_A*K_B_s_1*Rtot + 2*K_A*K_B_s_2*Ltot + 2*K_A*K_B_s_2*Rtot + 2*K_A*K_B_s_3*Ltot + 2*K_A*K_B_s_3*Rtot + 2*K_A*K_B_s_4*Ltot + 2*K_A*K_B_s_4*Rtot + 2*K_A*K_B_s_5*Ltot + 2*K_A*K_B_s_5*Rtot + 2*K_A*Ltot + 2*K_A*Rtot + K_B_s_1^2 + 2*K_B_s_1*K_B_s_2 + 2*K_B_s_1*K_B_s_3 + 2*K_B_s_1*K_B_s_4 + 2*K_B_s_1*K_B_s_5 + 2*K_B_s_1 + K_B_s_2^2 + 2*K_B_s_2*K_B_s_3 + 2*K_B_s_2*K_B_s_4 + 2*K_B_s_2*K_B_s_5 + 2*K_B_s_2 + K_B_s_3^2 + 2*K_B_s_3*K_B_s_4 + 2*K_B_s_3*K_B_s_5 + 2*K_B_s_3 + K_B_s_4^2 + 2*K_B_s_4*K_B_s_5 + 2*K_B_s_4 + K_B_s_5^2 + 2*K_B_s_5 + 1)^(1/2) + K_A*Ltot <> K_B_s_1 + K_B_s_2 + K_B_s_3 + K_B_s_4 + K_B_s_5 + K_A*Rtot + 1 and not -(K_B_s_1 + K_B_s_2 + K_B_s_3 + K_B_s_4 + K_B_s_5 - (K_A^2*Ltot^2 - 2*K_A^2*Ltot*Rtot + K_A^2*Rtot^2 + 2*K_A*K_B_s_1*Rtot + 2*K_A*K_B_s_2*Rtot + 2*K_A*K_B_s_3*Rtot + 2*K_A*K_B_s_4*Rtot + 2*K_A*K_B_s_5*Rtot + 2*K_A*Rtot + K_B_s_1^2 + 2*K_B_s_1*K_B_s_2 + 2*K_B_s_1*K_B_s_3 + 2*K_B_s_1*K_B_s_4 + 2*K_B_s_1*K_B_s_5 + 2*K_B_s_1 + K_B_s_2^2 + 2*K_B_s_2*K_B_s_3 + 2*K_B_s_2*K_B_s_4 + 2*K_B_s_2*K_B_s_5 + 2*K_B_s_2 + K_B_s_3^2 + 2*K_B_s_3*K_B_s_4 + 2*K_B_s_3*K_B_s_5 + 2*K_B_s_3 + K_B_s_4^2 + 2*K_B_s_4*K_B_s_5 + 2*K_B_s_4 + K_B_s_5^2 + 2*K_B_s_5 + 1)^(1/2) - K_A*Ltot + K_A*Rtot + 1)/(2*K_A) < Ltot and K_B_s_1 + K_B_s_2 + K_B_s_3 + K_B_s_4 + K_B_s_5 + (K_A^2*Ltot^2 - 2*K_A^2*Ltot*Rtot + K_A^2*Rtot^2 + 2*K_A*K_B_s_1*Ltot + 2*K_A*K_B_s_1*Rtot + 2*K_A*K_B_s_2*Ltot + 2*K_A*K_B_s_2*Rtot + 2*K_A*K_B_s_3*Ltot + 2*K_A*K_B_s_3*Rtot + 2*K_A*K_B_s_4*Ltot + 2*K_A*K_B_s_4*Rtot + 2*K_A*K_B_s_5*Ltot + 2*K_A*K_B_s_5*Rtot + 2*K_A*Ltot + 2*K_A*Rtot + K_B_s_1^2 + 2*K_B_s_1*K_B_s_2 + 2*K_B_s_1*K_B_s_3 + 2*K_B_s_1*K_B_s_4 + 2*K_B_s_1*K_B_s_5 + 2*K_B_s_1 + K_B_s_2^2 + 2*K_B_s_2*K_B_s_3 + 2*K_B_s_2*K_B_s_4 + 2*K_B_s_2*K_B_s_5 + 2*K_B_s_2 + K_B_s_3^2 + 2*K_B_s_3*K_B_s_4 + 2*K_B_s_3*K_B_s_5 + 2*K_B_s_3 + K_B_s_4^2 + 2*K_B_s_4*K_B_s_5 + 2*K_B_s_4 + K_B_s_5^2 + 2*K_B_s_5 + 1)^(1/2) + K_A*Rtot + 1 = K_A*Ltot or not -(K_B_s_1 + K_B_s_2 + K_B_s_3 + K_B_s_4 + K_B_s_5 + (K_A^2*Ltot^2 - 2*K_A^2*Ltot*Rtot + K_A^2*Rtot^2 + 2*K_A*K_B_s_1*Rtot + 2*K_A*K_B_s_2*Rtot + 2*K_A*K_B_s_3*Rtot + 2*K_A*K_B_s_4*Rtot + 2*K_A*K_B_s_5*Rtot + 2*K_A*Rtot + K_B_s_1^2 + 2*K_B_s_1*K_B_s_2 + 2*K_B_s_1*K_B_s_3 + 2*K_B_s_1*K_B_s_4 + 2*K_B_s_1*K_B_s_5 + 2*K_B_s_1 + K_B_s_2^2 + 2*K_B_s_2*K_B_s_3 + 2*K_B_s_2*K_B_s_4 + 2*K_B_s_2*K_B_s_5 + 2*K_B_s_2 + K_B_s_3^2 + 2*K_B_s_3*K_B_s_4 + 2*K_B_s_3*K_B_s_5 + 2*K_B_s_3 + K_B_s_4^2 + 2*K_B_s_4*K_B_s_5 + 2*K_B_s_4 + K_B_s_5^2 + 2*K_B_s_5 + 1)^(1/2) - K_A*Ltot + K_A*Rtot + 1)/(2*K_A) < Ltot and (K_A^2*Ltot^2 - 2*K_A^2*Ltot*Rtot + K_A^2*Rtot^2 + 2*K_A*K_B_s_1*Ltot + 2*K_A*K_B_s_1*Rtot + 2*K_A*K_B_s_2*Ltot + 2*K_A*K_B_s_2*Rtot + 2*K_A*K_B_s_3*Ltot + 2*K_A*K_B_s_3*Rtot + 2*K_A*K_B_s_4*Ltot + 2*K_A*K_B_s_4*Rtot + 2*K_A*K_B_s_5*Ltot + 2*K_A*K_B_s_5*Rtot + 2*K_A*Ltot + 2*K_A*Rtot + K_B_s_1^2 + 2*K_B_s_1*K_B_s_2 + 2*K_B_s_1*K_B_s_3 + 2*K_B_s_1*K_B_s_4 + 2*K_B_s_1*K_B_s_5 + 2*K_B_s_1 + K_B_s_2^2 + 2*K_B_s_2*K_B_s_3 + 2*K_B_s_2*K_B_s_4 + 2*K_B_s_2*K_B_s_5 + 2*K_B_s_2 + K_B_s_3^2 + 2*K_B_s_3*K_B_s_4 + 2*K_B_s_3*K_B_s_5 + 2*K_B_s_3 + K_B_s_4^2 + 2*K_B_s_4*K_B_s_5 + 2*K_B_s_4 + K_B_s_5^2 + 2*K_B_s_5 + 1)^(1/2) + K_A*Ltot = K_B_s_1 + K_B_s_2 + K_B_s_3 + K_B_s_4 + K_B_s_5 + K_A*Rtot + 1 and K_B_s_1 + K_B_s_2 + K_B_s_3 + K_B_s_4 + K_B_s_5 + (K_A^2*Ltot^2 - 2*K_A^2*Ltot*Rtot + K_A^2*Rtot^2 + 2*K_A*K_B_s_1*Rtot + 2*K_A*K_B_s_2*Rtot + 2*K_A*K_B_s_3*Rtot + 2*K_A*K_B_s_4*Rtot + 2*K_A*K_B_s_5*Rtot + 2*K_A*Rtot + K_B_s_1^2 + 2*K_B_s_1*K_B_s_2 + 2*K_B_s_1*K_B_s_3 + 2*K_B_s_1*K_B_s_4 + 2*K_B_s_1*K_B_s_5 + 2*K_B_s_1 + K_B_s_2^2 + 2*K_B_s_2*K_B_s_3 + 2*K_B_s_2*K_B_s_4 + 2*K_B_s_2*K_B_s_5 + 2*K_B_s_2 + K_B_s_3^2 + 2*K_B_s_3*K_B_s_4 + 2*K_B_s_3*K_B_s_5 + 2*K_B_s_3 + K_B_s_4^2 + 2*K_B_s_4*K_B_s_5 + 2*K_B_s_4 + K_B_s_5^2 + 2*K_B_s_5 + 1)^(1/2) + K_A*Rtot + 1 < K_A*Ltot and K_B_s_1 + K_B_s_2 + K_B_s_3 + K_B_s_4 + K_B_s_5 + (K_A^2*Ltot^2 - 2*K_A^2*Ltot*Rtot + K_A^2*Rtot^2 + 2*K_A*K_B_s_1*Ltot + 2*K_A*K_B_s_1*Rtot + 2*K_A*K_B_s_2*Ltot + 2*K_A*K_B_s_2*Rtot + 2*K_A*K_B_s_3*Ltot + 2*K_A*K_B_s_3*Rtot + 2*K_A*K_B_s_4*Ltot + 2*K_A*K_B_s_4*Rtot + 2*K_A*K_B_s_5*Ltot + 2*K_A*K_B_s_5*Rtot + 2*K_A*Ltot + 2*K_A*Rtot + K_B_s_1^2 + 2*K_B_s_1*K_B_s_2 + 2*K_B_s_1*K_B_s_3 + 2*K_B_s_1*K_B_s_4 + 2*K_B_s_1*K_B_s_5 + 2*K_B_s_1 + K_B_s_2^2 + 2*K_B_s_2*K_B_s_3 + 2*K_B_s_2*K_B_s_4 + 2*K_B_s_2*K_B_s_5 + 2*K_B_s_2 + K_B_s_3^2 + 2*K_B_s_3*K_B_s_4 + 2*K_B_s_3*K_B_s_5 + 2*K_B_s_3 + K_B_s_4^2 + 2*K_B_s_4*K_B_s_5 + 2*K_B_s_4 + K_B_s_5^2 + 2*K_B_s_5 + 1)^(1/2) + K_A*Rtot + 1 <> K_A*Ltot or (not K_B_s_1 + K_B_s_2 + K_B_s_3 + K_B_s_4 + K_B_s_5 - (K_A^2*Ltot^2 - 2*K_A^2*Ltot*Rtot + K_A^2*Rtot^2 + 2*K_A*K_B_s_1*Ltot + 2*K_A*K_B_s_1*Rtot + 2*K_A*K_B_s_2*Ltot + 2*K_A*K_B_s_2*Rtot + 2*K_A*K_B_s_3*Ltot + 2*K_A*K_B_s_3*Rtot + 2*K_A*K_B_s_4*Ltot + 2*K_A*K_B_s_4*Rtot + 2*K_A*K_B_s_5*Ltot + 2*K_A*K_B_s_5*Rtot + 2*K_A*Ltot + 2*K_A*Rtot + K_B_s_1^2 + 2*K_B_s_1*K_B_s_2 + 2*K_B_s_1*K_B_s_3 + 2*K_B_s_1*K_B_s_4 + 2*K_B_s_1*K_B_s_5 + 2*K_B_s_1 + K_B_s_2^2 + 2*K_B_s_2*K_B_s_3 + 2*K_B_s_2*K_B_s_4 + 2*K_B_s_2*K_B_s_5 + 2*K_B_s_2 + K_B_s_3^2 + 2*K_B_s_3*K_B_s_4 + 2*K_B_s_3*K_B_s_5 + 2*K_B_s_3 + K_B_s_4^2 + 2*K_B_s_4*K_B_s_5 + 2*K_B_s_4 + K_B_s_5^2 + 2*K_B_s_5 + 1)^(1/2) + K_A*Rtot + 1 < K_A*Ltot or not -(K_B_s_1 + K_B_s_2 + K_B_s_3 + K_B_s_4 + K_B_s_5 - (K_A^2*Ltot^2 - 2*K_A^2*Ltot*Rtot + K_A^2*Rtot^2 + 2*K_A*K_B_s_1*Ltot + 2*K_A*K_B_s_1*Rtot + 2*K_A*K_B_s_2*Ltot + 2*K_A*K_B_s_2*Rtot + 2*K_A*K_B_s_3*Ltot + 2*K_A*K_B_s_3*Rtot + 2*K_A*K_B_s_4*Ltot + 2*K_A*K_B_s_4*Rtot + 2*K_A*K_B_s_5*Ltot + 2*K_A*K_B_s_5*Rtot + 2*K_A*Ltot + 2*K_A*Rtot + K_B_s_1^2 + 2*K_B_s_1*K_B_s_2 + 2*K_B_s_1*K_B_s_3 + 2*K_B_s_1*K_B_s_4 + 2*K_B_s_1*K_B_s_5 + 2*K_B_s_1 + K_B_s_2^2 + 2*K_B_s_2*K_B_s_3 + 2*K_B_s_2*K_B_s_4 + 2*K_B_s_2*K_B_s_5 + 2*K_B_s_2 + K_B_s_3^2 + 2*K_B_s_3*K_B_s_4 + 2*K_B_s_3*K_B_s_5 + 2*K_B_s_3 + K_B_s_4^2 + 2*K_B_s_4*K_B_s_5 + 2*K_B_s_4 + K_B_s_5^2 + 2*K_B_s_5 + 1)^(1/2) - K_A*Ltot + K_A*Rtot + 1)/(2*K_A) < Ltot) and -(K_B_s_1 + K_B_s_2 + K_B_s_3 + K_B_s_4 + K_B_s_5 + (K_A^2*Ltot^2 - 2*K_A^2*Ltot*Rtot + K_A^2*Rtot^2 + 2*K_A*K_B_s_1*Ltot + 2*K_A*K_B_s_1*Rtot + 2*K_A*K_B_s_2*Ltot + 2*K_A*K_B_s_2*Rtot + 2*K_A*K_B_s_3*Ltot + 2*K_A*K_B_s_3*Rtot + 2*K_A*K_B_s_4*Ltot + 2*K_A*K_B_s_4*Rtot + 2*K_A*K_B_s_5*Ltot + 2*K_A*K_B_s_5*Rtot + 2*K_A*Ltot + 2*K_A*Rtot + K_B_s_1^2 + 2*K_B_s_1*K_B_s_2 + 2*K_B_s_1*K_B_s_3 + 2*K_B_s_1*K_B_s_4 + 2*K_B_s_1*K_B_s_5 + 2*K_B_s_1 + K_B_s_2^2 + 2*K_B_s_2*K_B_s_3 + 2*K_B_s_2*K_B_s_4 + 2*K_B_s_2*K_B_s_5 + 2*K_B_s_2 + K_B_s_3^2 + 2*K_B_s_3*K_B_s_4 + 2*K_B_s_3*K_B_s_5 + 2*K_B_s_3 + K_B_s_4^2 + 2*K_B_s_4*K_B_s_5 + 2*K_B_s_4 + K_B_s_5^2 + 2*K_B_s_5 + 1)^(1/2) - K_A*Ltot + K_A*Rtot + 1)/(2*K_A) < Ltot and (K_A^2*Ltot^2 - 2*K_A^2*Ltot*Rtot + K_A^2*Rtot^2 + 2*K_A*K_B_s_1*Ltot + 2*K_A*K_B_s_1*Rtot + 2*K_A*K_B_s_2*Ltot + 2*K_A*K_B_s_2*Rtot + 2*K_A*K_B_s_3*Ltot + 2*K_A*K_B_s_3*Rtot + 2*K_A*K_B_s_4*Ltot + 2*K_A*K_B_s_4*Rtot + 2*K_A*K_B_s_5*Ltot + 2*K_A*K_B_s_5*Rtot + 2*K_A*Ltot + 2*K_A*Rtot + K_B_s_1^2 + 2*K_B_s_1*K_B_s_2 + 2*K_B_s_1*K_B_s_3 + 2*K_B_s_1*K_B_s_4 + 2*K_B_s_1*K_B_s_5 + 2*K_B_s_1 + K_B_s_2^2 + 2*K_B_s_2*K_B_s_3 + 2*K_B_s_2*K_B_s_4 + 2*K_B_s_2*K_B_s_5 + 2*K_B_s_2 + K_B_s_3^2 + 2*K_B_s_3*K_B_s_4 + 2*K_B_s_3*K_B_s_5 + 2*K_B_s_3 + K_B_s_4^2 + 2*K_B_s_4*K_B_s_5 + 2*K_B_s_4 + K_B_s_5^2 + 2*K_B_s_5 + 1)^(1/2) + K_A*Ltot <> K_B_s_1 + K_B_s_2 + K_B_s_3 + K_B_s_4 + K_B_s_5 + K_A*Rtot + 1 and K_A^2*Ltot^2 + K_A^2*Rtot^2 + 2*K_A*K_B_s_1*Ltot + 2*K_A*K_B_s_1*Rtot + 2*K_A*K_B_s_2*Ltot + 2*K_A*K_B_s_2*Rtot + 2*K_A*K_B_s_3*Ltot + 2*K_A*K_B_s_3*Rtot + 2*K_A*K_B_s_4*Ltot + 2*K_A*K_B_s_4*Rtot + 2*K_A*K_B_s_5*Ltot + 2*K_A*K_B_s_5*Rtot + 2*K_A*Ltot + 2*K_A*Rtot + K_B_s_1^2 + 2*K_B_s_1*K_B_s_2 + 2*K_B_s_1*K_B_s_3 + 2*K_B_s_1*K_B_s_4 + 2*K_B_s_1*K_B_s_5 + 2*K_B_s_1 + K_B_s_2^2 + 2*K_B_s_2*K_B_s_3 + 2*K_B_s_2*K_B_s_4 + 2*K_B_s_2*K_B_s_5 + 2*K_B_s_2 + K_B_s_3^2 + 2*K_B_s_3*K_B_s_4 + 2*K_B_s_3*K_B_s_5 + 2*K_B_s_3 + K_B_s_4^2 + 2*K_B_s_4*K_B_s_5 + 2*K_B_s_4 + K_B_s_5^2 + 2*K_B_s_5 + 1 < 2*K_A^2*Ltot*Rtot and K_B_s_1 + K_B_s_2 + K_B_s_3 + K_B_s_4 + K_B_s_5 + (K_A^2*Ltot^2 - 2*K_A^2*Ltot*Rtot + K_A^2*Rtot^2 + 2*K_A*K_B_s_1*Ltot + 2*K_A*K_B_s_1*Rtot + 2*K_A*K_B_s_2*Ltot + 2*K_A*K_B_s_2*Rtot + 2*K_A*K_B_s_3*Ltot + 2*K_A*K_B_s_3*Rtot + 2*K_A*K_B_s_4*Ltot + 2*K_A*K_B_s_4*Rtot + 2*K_A*K_B_s_5*Ltot + 2*K_A*K_B_s_5*Rtot + 2*K_A*Ltot + 2*K_A*Rtot + K_B_s_1^2 + 2*K_B_s_1*K_B_s_2 + 2*K_B_s_1*K_B_s_3 + 2*K_B_s_1*K_B_s_4 + 2*K_B_s_1*K_B_s_5 + 2*K_B_s_1 + K_B_s_2^2 + 2*K_B_s_2*K_B_s_3 + 2*K_B_s_2*K_B_s_4 + 2*K_B_s_2*K_B_s_5 + 2*K_B_s_2 + K_B_s_3^2 + 2*K_B_s_3*K_B_s_4 + 2*K_B_s_3*K_B_s_5 + 2*K_B_s_3 + K_B_s_4^2 + 2*K_B_s_4*K_B_s_5 + 2*K_B_s_4 + K_B_s_5^2 + 2*K_B_s_5 + 1)^(1/2) + K_A*Rtot + 1 < K_A*Ltot or K_B_s_1 + K_B_s_2 + K_B_s_3 + K_B_s_4 + K_B_s_5 - (K_A^2*Ltot^2 - 2*K_A^2*Ltot*Rtot + K_A^2*Rtot^2 + 2*K_A*K_B_s_1*Ltot + 2*K_A*K_B_s_1*Rtot + 2*K_A*K_B_s_2*Ltot + 2*K_A*K_B_s_2*Rtot + 2*K_A*K_B_s_3*Ltot + 2*K_A*K_B_s_3*Rtot + 2*K_A*K_B_s_4*Ltot + 2*K_A*K_B_s_4*Rtot + 2*K_A*K_B_s_5*Ltot + 2*K_A*K_B_s_5*Rtot + 2*K_A*Ltot + 2*K_A*Rtot + K_B_s_1^2 + 2*K_B_s_1*K_B_s_2 + 2*K_B_s_1*K_B_s_3 + 2*K_B_s_1*K_B_s_4 + 2*K_B_s_1*K_B_s_5 + 2*K_B_s_1 + K_B_s_2^2 + 2*K_B_s_2*K_B_s_3 + 2*K_B_s_2*K_B_s_4 + 2*K_B_s_2*K_B_s_5 + 2*K_B_s_2 + K_B_s_3^2 + 2*K_B_s_3*K_B_s_4 + 2*K_B_s_3*K_B_s_5 + 2*K_B_s_3 + K_B_s_4^2 + 2*K_B_s_4*K_B_s_5 + 2*K_B_s_4 + K_B_s_5^2 + 2*K_B_s_5 + 1)^(1/2) + K_A*Rtot + 1 < K_A*Ltot and (K_A^2*Ltot^2 - 2*K_A^2*Ltot*Rtot + K_A^2*Rtot^2 + 2*K_A*K_B_s_1*Ltot + 2*K_A*K_B_s_1*Rtot + 2*K_A*K_B_s_2*Ltot + 2*K_A*K_B_s_2*Rtot + 2*K_A*K_B_s_3*Ltot + 2*K_A*K_B_s_3*Rtot + 2*K_A*K_B_s_4*Ltot + 2*K_A*K_B_s_4*Rtot + 2*K_A*K_B_s_5*Ltot + 2*K_A*K_B_s_5*Rtot + 2*K_A*Ltot + 2*K_A*Rtot + K_B_s_1^2 + 2*K_B_s_1*K_B_s_2 + 2*K_B_s_1*K_B_s_3 + 2*K_B_s_1*K_B_s_4 + 2*K_B_s_1*K_B_s_5 + 2*K_B_s_1 + K_B_s_2^2 + 2*K_B_s_2*K_B_s_3 + 2*K_B_s_2*K_B_s_4 + 2*K_B_s_2*K_B_s_5 + 2*K_B_s_2 + K_B_s_3^2 + 2*K_B_s_3*K_B_s_4 + 2*K_B_s_3*K_B_s_5 + 2*K_B_s_3 + K_B_s_4^2 + 2*K_B_s_4*K_B_s_5 + 2*K_B_s_4 + K_B_s_5^2 + 2*K_B_s_5 + 1)^(1/2) + K_A*Ltot <> K_B_s_1 + K_B_s_2 + K_B_s_3 + K_B_s_4 + K_B_s_5 + K_A*Rtot + 1 and -(K_B_s_1 + K_B_s_2 + K_B_s_3 + K_B_s_4 + K_B_s_5 - (K_A^2*Ltot^2 - 2*K_A^2*Ltot*Rtot + K_A^2*Rtot^2 + 2*K_A*K_B_s_1*Ltot + 2*K_A*K_B_s_1*Rtot + 2*K_A*K_B_s_2*Ltot + 2*K_A*K_B_s_2*Rtot + 2*K_A*K_B_s_3*Ltot + 2*K_A*K_B_s_3*Rtot + 2*K_A*K_B_s_4*Ltot + 2*K_A*K_B_s_4*Rtot + 2*K_A*K_B_s_5*Ltot + 2*K_A*K_B_s_5*Rtot + 2*K_A*Ltot + 2*K_A*Rtot + K_B_s_1^2 + 2*K_B_s_1*K_B_s_2 + 2*K_B_s_1*K_B_s_3 + 2*K_B_s_1*K_B_s_4 + 2*K_B_s_1*K_B_s_5 + 2*K_B_s_1 + K_B_s_2^2 + 2*K_B_s_2*K_B_s_3 + 2*K_B_s_2*K_B_s_4 + 2*K_B_s_2*K_B_s_5 + 2*K_B_s_2 + K_B_s_3^2 + 2*K_B_s_3*K_B_s_4 + 2*K_B_s_3*K_B_s_5 + 2*K_B_s_3 + K_B_s_4^2 + 2*K_B_s_4*K_B_s_5 + 2*K_B_s_4 + K_B_s_5^2 + 2*K_B_s_5 + 1)^(1/2) - K_A*Ltot + K_A*Rtot + 1)/(2*K_A) < Ltot and (not -(K_B_s_1 + K_B_s_2 + K_B_s_3 + K_B_s_4 + K_B_s_5 + (K_A^2*Ltot^2 - 2*K_A^2*Ltot*Rtot + K_A^2*Rtot^2 + 2*K_A*K_B_s_1*Ltot + 2*K_A*K_B_s_1*Rtot + 2*K_A*K_B_s_2*Ltot + 2*K_A*K_B_s_2*Rtot + 2*K_A*K_B_s_3*Ltot + 2*K_A*K_B_s_3*Rtot + 2*K_A*K_B_s_4*Ltot + 2*K_A*K_B_s_4*Rtot + 2*K_A*K_B_s_5*Ltot + 2*K_A*K_B_s_5*Rtot + 2*K_A*Ltot + 2*K_A*Rtot + K_B_s_1^2 + 2*K_B_s_1*K_B_s_2 + 2*K_B_s_1*K_B_s_3 + 2*K_B_s_1*K_B_s_4 + 2*K_B_s_1*K_B_s_5 + 2*K_B_s_1 + K_B_s_2^2 + 2*K_B_s_2*K_B_s_3 + 2*K_B_s_2*K_B_s_4 + 2*K_B_s_2*K_B_s_5 + 2*K_B_s_2 + K_B_s_3^2 + 2*K_B_s_3*K_B_s_4 + 2*K_B_s_3*K_B_s_5 + 2*K_B_s_3 + K_B_s_4^2 + 2*K_B_s_4*K_B_s_5 + 2*K_B_s_4 + K_B_s_5^2 + 2*K_B_s_5 + 1)^(1/2) - K_A*Ltot + K_A*Rtot + 1)/(2*K_A) < Ltot and K_B_s_1 + K_B_s_2 + K_B_s_3 + K_B_s_4 + K_B_s_5 + (K_A^2*Ltot^2 - 2*K_A^2*Ltot*Rtot + K_A^2*Rtot^2 + 2*K_A*K_B_s_1*Ltot + 2*K_A*K_B_s_1*Rtot + 2*K_A*K_B_s_2*Ltot + 2*K_A*K_B_s_2*Rtot + 2*K_A*K_B_s_3*Ltot + 2*K_A*K_B_s_3*Rtot + 2*K_A*K_B_s_4*Ltot + 2*K_A*K_B_s_4*Rtot + 2*K_A*K_B_s_5*Ltot + 2*K_A*K_B_s_5*Rtot + 2*K_A*Ltot + 2*K_A*Rtot + K_B_s_1^2 + 2*K_B_s_1*K_B_s_2 + 2*K_B_s_1*K_B_s_3 + 2*K_B_s_1*K_B_s_4 + 2*K_B_s_1*K_B_s_5 + 2*K_B_s_1 + K_B_s_2^2 + 2*K_B_s_2*K_B_s_3 + 2*K_B_s_2*K_B_s_4 + 2*K_B_s_2*K_B_s_5 + 2*K_B_s_2 + K_B_s_3^2 + 2*K_B_s_3*K_B_s_4 + 2*K_B_s_3*K_B_s_5 + 2*K_B_s_3 + K_B_s_4^2 + 2*K_B_s_4*K_B_s_5 + 2*K_B_s_4 + K_B_s_5^2 + 2*K_B_s_5 + 1)^(1/2) + K_A*Rtot + 1 < K_A*Ltot or not K_B_s_1 + K_B_s_2 + K_B_s_3 + K_B_s_4 + K_B_s_5 + (K_A^2*Ltot^2 - 2*K_A^2*Ltot*Rtot + K_A^2*Rtot^2 + 2*K_A*K_B_s_1*Ltot + 2*K_A*K_B_s_1*Rtot + 2*K_A*K_B_s_2*Ltot + 2*K_A*K_B_s_2*Rtot + 2*K_A*K_B_s_3*Ltot + 2*K_A*K_B_s_3*Rtot + 2*K_A*K_B_s_4*Ltot + 2*K_A*K_B_s_4*Rtot + 2*K_A*K_B_s_5*Ltot + 2*K_A*K_B_s_5*Rtot + 2*K_A*Ltot + 2*K_A*Rtot + K_B_s_1^2 + 2*K_B_s_1*K_B_s_2 + 2*K_B_s_1*K_B_s_3 + 2*K_B_s_1*K_B_s_4 + 2*K_B_s_1*K_B_s_5 + 2*K_B_s_1 + K_B_s_2^2 + 2*K_B_s_2*K_B_s_3 + 2*K_B_s_2*K_B_s_4 + 2*K_B_s_2*K_B_s_5 + 2*K_B_s_2 + K_B_s_3^2 + 2*K_B_s_3*K_B_s_4 + 2*K_B_s_3*K_B_s_5 + 2*K_B_s_3 + K_B_s_4^2 + 2*K_B_s_4*K_B_s_5 + 2*K_B_s_4 + K_B_s_5^2 + 2*K_B_s_5 + 1)^(1/2) + K_A*Rtot + 1 <= K_A*Ltot) and K_A^2*Ltot^2 + K_A^2*Rtot^2 + 2*K_A*K_B_s_1*Ltot + 2*K_A*K_B_s_1*Rtot + 2*K_A*K_B_s_2*Ltot + 2*K_A*K_B_s_2*Rtot + 2*K_A*K_B_s_3*Ltot + 2*K_A*K_B_s_3*Rtot + 2*K_A*K_B_s_4*Ltot + 2*K_A*K_B_s_4*Rtot + 2*K_A*K_B_s_5*Ltot + 2*K_A*K_B_s_5*Rtot + 2*K_A*Ltot + 2*K_A*Rtot + K_B_s_1^2 + 2*K_B_s_1*K_B_s_2 + 2*K_B_s_1*K_B_s_3 + 2*K_B_s_1*K_B_s_4 + 2*K_B_s_1*K_B_s_5 + 2*K_B_s_1 + K_B_s_2^2 + 2*K_B_s_2*K_B_s_3 + 2*K_B_s_2*K_B_s_4 + 2*K_B_s_2*K_B_s_5 + 2*K_B_s_2 + K_B_s_3^2 + 2*K_B_s_3*K_B_s_4 + 2*K_B_s_3*K_B_s_5 + 2*K_B_s_3 + K_B_s_4^2 + 2*K_B_s_4*K_B_s_5 + 2*K_B_s_4 + K_B_s_5^2 + 2*K_B_s_5 + 1 < 2*K_A^2*Ltot*Rtot or not -(K_B_s_1 + K_B_s_2 + K_B_s_3 + K_B_s_4 + K_B_s_5 + (K_A^2*Ltot^2 - 2*K_A^2*Ltot*Rtot + K_A^2*Rtot^2 + 2*K_A*K_B_s_1*Ltot + 2*K_A*K_B_s_1*Rtot + 2*K_A*K_B_s_2*Ltot + 2*K_A*K_B_s_2*Rtot + 2*K_A*K_B_s_3*Ltot + 2*K_A*K_B_s_3*Rtot + 2*K_A*K_B_s_4*Ltot + 2*K_A*K_B_s_4*Rtot + 2*K_A*K_B_s_5*Ltot + 2*K_A*K_B_s_5*Rtot + 2*K_A*Ltot + 2*K_A*Rtot + K_B_s_1^2 + 2*K_B_s_1*K_B_s_2 + 2*K_B_s_1*K_B_s_3 + 2*K_B_s_1*K_B_s_4 + 2*K_B_s_1*K_B_s_5 + 2*K_B_s_1 + K_B_s_2^2 + 2*K_B_s_2*K_B_s_3 + 2*K_B_s_2*K_B_s_4 + 2*K_B_s_2*K_B_s_5 + 2*K_B_s_2 + K_B_s_3^2 + 2*K_B_s_3*K_B_s_4 + 2*K_B_s_3*K_B_s_5 + 2*K_B_s_3 + K_B_s_4^2 + 2*K_B_s_4*K_B_s_5 + 2*K_B_s_4 + K_B_s_5^2 + 2*K_B_s_5 + 1)^(1/2) - K_A*Ltot + K_A*Rtot + 1)/(2*K_A) < Ltot and K_B_s_1 + K_B_s_2 + K_B_s_3 + K_B_s_4 + K_B_s_5 - (K_A^2*Ltot^2 - 2*K_A^2*Ltot*Rtot + K_A^2*Rtot^2 + 2*K_A*K_B_s_1*Ltot + 2*K_A*K_B_s_1*Rtot + 2*K_A*K_B_s_2*Ltot + 2*K_A*K_B_s_2*Rtot + 2*K_A*K_B_s_3*Ltot + 2*K_A*K_B_s_3*Rtot + 2*K_A*K_B_s_4*Ltot + 2*K_A*K_B_s_4*Rtot + 2*K_A*K_B_s_5*Ltot + 2*K_A*K_B_s_5*Rtot + 2*K_A*Ltot + 2*K_A*Rtot + K_B_s_1^2 + 2*K_B_s_1*K_B_s_2 + 2*K_B_s_1*K_B_s_3 + 2*K_B_s_1*K_B_s_4 + 2*K_B_s_1*K_B_s_5 + 2*K_B_s_1 + K_B_s_2^2 + 2*K_B_s_2*K_B_s_3 + 2*K_B_s_2*K_B_s_4 + 2*K_B_s_2*K_B_s_5 + 2*K_B_s_2 + K_B_s_3^2 + 2*K_B_s_3*K_B_s_4 + 2*K_B_s_3*K_B_s_5 + 2*K_B_s_3 + K_B_s_4^2 + 2*K_B_s_4*K_B_s_5 + 2*K_B_s_4 + K_B_s_5^2 + 2*K_B_s_5 + 1)^(1/2) + K_A*Rtot + 1 < K_A*Ltot and (K_A^2*Ltot^2 - 2*K_A^2*Ltot*Rtot + K_A^2*Rtot^2 + 2*K_A*K_B_s_1*Ltot + 2*K_A*K_B_s_1*Rtot + 2*K_A*K_B_s_2*Ltot + 2*K_A*K_B_s_2*Rtot + 2*K_A*K_B_s_3*Ltot + 2*K_A*K_B_s_3*Rtot + 2*K_A*K_B_s_4*Ltot + 2*K_A*K_B_s_4*Rtot + 2*K_A*K_B_s_5*Ltot + 2*K_A*K_B_s_5*Rtot + 2*K_A*Ltot + 2*K_A*Rtot + K_B_s_1^2 + 2*K_B_s_1*K_B_s_2 + 2*K_B_s_1*K_B_s_3 + 2*K_B_s_1*K_B_s_4 + 2*K_B_s_1*K_B_s_5 + 2*K_B_s_1 + K_B_s_2^2 + 2*K_B_s_2*K_B_s_3 + 2*K_B_s_2*K_B_s_4 + 2*K_B_s_2*K_B_s_5 + 2*K_B_s_2 + K_B_s_3^2 + 2*K_B_s_3*K_B_s_4 + 2*K_B_s_3*K_B_s_5 + 2*K_B_s_3 + K_B_s_4^2 + 2*K_B_s_4*K_B_s_5 + 2*K_B_s_4 + K_B_s_5^2 + 2*K_B_s_5 + 1)^(1/2) + K_A*Ltot <> K_B_s_1 + K_B_s_2 + K_B_s_3 + K_B_s_4 + K_B_s_5 + K_A*Rtot + 1 and not -(K_B_s_1 + K_B_s_2 + K_B_s_3 + K_B_s_4 + K_B_s_5 - (K_A^2*Ltot^2 - 2*K_A^2*Ltot*Rtot + K_A^2*Rtot^2 + 2*K_A*K_B_s_1*Ltot + 2*K_A*K_B_s_1*Rtot + 2*K_A*K_B_s_2*Ltot + 2*K_A*K_B_s_2*Rtot + 2*K_A*K_B_s_3*Ltot + 2*K_A*K_B_s_3*Rtot + 2*K_A*K_B_s_4*Ltot + 2*K_A*K_B_s_4*Rtot + 2*K_A*K_B_s_5*Ltot + 2*K_A*K_B_s_5*Rtot + 2*K_A*Ltot + 2*K_A*Rtot + K_B_s_1^2 + 2*K_B_s_1*K_B_s_2 + 2*K_B_s_1*K_B_s_3 + 2*K_B_s_1*K_B_s_4 + 2*K_B_s_1*K_B_s_5 + 2*K_B_s_1 + K_B_s_2^2 + 2*K_B_s_2*K_B_s_3 + 2*K_B_s_2*K_B_s_4 + 2*K_B_s_2*K_B_s_5 + 2*K_B_s_2 + K_B_s_3^2 + 2*K_B_s_3*K_B_s_4 + 2*K_B_s_3*K_B_s_5 + 2*K_B_s_3 + K_B_s_4^2 + 2*K_B_s_4*K_B_s_5 + 2*K_B_s_4 + K_B_s_5^2 + 2*K_B_s_5 + 1)^(1/2) - K_A*Ltot + K_A*Rtot + 1)/(2*K_A) < Ltot and K_B_s_1 + K_B_s_2 + K_B_s_3 + K_B_s_4 + K_B_s_5 + (K_A^2*Ltot^2 - 2*K_A^2*Ltot*Rtot + K_A^2*Rtot^2 + 2*K_A*K_B_s_1*Ltot + 2*K_A*K_B_s_1*Rtot + 2*K_A*K_B_s_2*Ltot + 2*K_A*K_B_s_2*Rtot + 2*K_A*K_B_s_3*Ltot + 2*K_A*K_B_s_3*Rtot + 2*K_A*K_B_s_4*Ltot + 2*K_A*K_B_s_4*Rtot + 2*K_A*K_B_s_5*Ltot + 2*K_A*K_B_s_5*Rtot + 2*K_A*Ltot + 2*K_A*Rtot + K_B_s_1^2 + 2*K_B_s_1*K_B_s_2 + 2*K_B_s_1*K_B_s_3 + 2*K_B_s_1*K_B_s_4 + 2*K_B_s_1*K_B_s_5 + 2*K_B_s_1 + K_B_s_2^2 + 2*K_B_s_2*K_B_s_3 + 2*K_B_s_2*K_B_s_4 + 2*K_B_s_2*K_B_s_5 + 2*K_B_s_2 + K_B_s_3^2 + 2*K_B_s_3*K_B_s_4 + 2*K_B_s_3*K_B_s_5 + 2*K_B_s_3 + K_B_s_4^2 + 2*K_B_s_4*K_B_s_5 + 2*K_B_s_4 + K_B_s_5^2 + 2*K_B_s_5 + 1)^(1/2) + K_A*Rtot + 1 < K_A*Ltot or -(K_B_s_1 + K_B_s_2 + K_B_s_3 + K_B_s_4 + K_B_s_5 + (K_A^2*Ltot^2 - 2*K_A^2*Ltot*Rtot + K_A^2*Rtot^2 + 2*K_A*K_B_s_1*Ltot + 2*K_A*K_B_s_1*Rtot + 2*K_A*K_B_s_2*Ltot + 2*K_A*K_B_s_2*Rtot + 2*K_A*K_B_s_3*Ltot + 2*K_A*K_B_s_3*Rtot + 2*K_A*K_B_s_4*Ltot + 2*K_A*K_B_s_4*Rtot + 2*K_A*K_B_s_5*Ltot + 2*K_A*K_B_s_5*Rtot + 2*K_A*Ltot + 2*K_A*Rtot + K_B_s_1^2 + 2*K_B_s_1*K_B_s_2 + 2*K_B_s_1*K_B_s_3 + 2*K_B_s_1*K_B_s_4 + 2*K_B_s_1*K_B_s_5 + 2*K_B_s_1 + K_B_s_2^2 + 2*K_B_s_2*K_B_s_3 + 2*K_B_s_2*K_B_s_4 + 2*K_B_s_2*K_B_s_5 + 2*K_B_s_2 + K_B_s_3^2 + 2*K_B_s_3*K_B_s_4 + 2*K_B_s_3*K_B_s_5 + 2*K_B_s_3 + K_B_s_4^2 + 2*K_B_s_4*K_B_s_5 + 2*K_B_s_4 + K_B_s_5^2 + 2*K_B_s_5 + 1)^(1/2) - K_A*Ltot + K_A*Rtot + 1)/(2*K_A) < Ltot and K_B_s_1 + K_B_s_2 + K_B_s_3 + K_B_s_4 + K_B_s_5 - (K_A^2*Ltot^2 - 2*K_A^2*Ltot*Rtot + K_A^2*Rtot^2 + 2*K_A*K_B_s_1*Ltot + 2*K_A*K_B_s_1*Rtot + 2*K_A*K_B_s_2*Ltot + 2*K_A*K_B_s_2*Rtot + 2*K_A*K_B_s_3*Ltot + 2*K_A*K_B_s_3*Rtot + 2*K_A*K_B_s_4*Ltot + 2*K_A*K_B_s_4*Rtot + 2*K_A*K_B_s_5*Ltot + 2*K_A*K_B_s_5*Rtot + 2*K_A*Ltot + 2*K_A*Rtot + K_B_s_1^2 + 2*K_B_s_1*K_B_s_2 + 2*K_B_s_1*K_B_s_3 + 2*K_B_s_1*K_B_s_4 + 2*K_B_s_1*K_B_s_5 + 2*K_B_s_1 + K_B_s_2^2 + 2*K_B_s_2*K_B_s_3 + 2*K_B_s_2*K_B_s_4 + 2*K_B_s_2*K_B_s_5 + 2*K_B_s_2 + K_B_s_3^2 + 2*K_B_s_3*K_B_s_4 + 2*K_B_s_3*K_B_s_5 + 2*K_B_s_3 + K_B_s_4^2 + 2*K_B_s_4*K_B_s_5 + 2*K_B_s_4 + K_B_s_5^2 + 2*K_B_s_5 + 1)^(1/2) + K_A*Rtot + 1 < K_A*Ltot and (K_A^2*Ltot^2 - 2*K_A^2*Ltot*Rtot + K_A^2*Rtot^2 + 2*K_A*K_B_s_1*Ltot + 2*K_A*K_B_s_1*Rtot + 2*K_A*K_B_s_2*Ltot + 2*K_A*K_B_s_2*Rtot + 2*K_A*K_B_s_3*Ltot + 2*K_A*K_B_s_3*Rtot + 2*K_A*K_B_s_4*Ltot + 2*K_A*K_B_s_4*Rtot + 2*K_A*K_B_s_5*Ltot + 2*K_A*K_B_s_5*Rtot + 2*K_A*Ltot + 2*K_A*Rtot + K_B_s_1^2 + 2*K_B_s_1*K_B_s_2 + 2*K_B_s_1*K_B_s_3 + 2*K_B_s_1*K_B_s_4 + 2*K_B_s_1*K_B_s_5 + 2*K_B_s_1 + K_B_s_2^2 + 2*K_B_s_2*K_B_s_3 + 2*K_B_s_2*K_B_s_4 + 2*K_B_s_2*K_B_s_5 + 2*K_B_s_2 + K_B_s_3^2 + 2*K_B_s_3*K_B_s_4 + 2*K_B_s_3*K_B_s_5 + 2*K_B_s_3 + K_B_s_4^2 + 2*K_B_s_4*K_B_s_5 + 2*K_B_s_4 + K_B_s_5^2 + 2*K_B_s_5 + 1)^(1/2) + K_A*Ltot <> K_B_s_1 + K_B_s_2 + K_B_s_3 + K_B_s_4 + K_B_s_5 + K_A*Rtot + 1 and -(K_B_s_1 + K_B_s_2 + K_B_s_3 + K_B_s_4 + K_B_s_5 - (K_A^2*Ltot^2 - 2*K_A^2*Ltot*Rtot + K_A^2*Rtot^2 + 2*K_A*K_B_s_1*Ltot + 2*K_A*K_B_s_1*Rtot + 2*K_A*K_B_s_2*Ltot + 2*K_A*K_B_s_2*Rtot + 2*K_A*K_B_s_3*Ltot + 2*K_A*K_B_s_3*Rtot + 2*K_A*K_B_s_4*Ltot + 2*K_A*K_B_s_4*Rtot + 2*K_A*K_B_s_5*Ltot + 2*K_A*K_B_s_5*Rtot + 2*K_A*Ltot + 2*K_A*Rtot + K_B_s_1^2 + 2*K_B_s_1*K_B_s_2 + 2*K_B_s_1*K_B_s_3 + 2*K_B_s_1*K_B_s_4 + 2*K_B_s_1*K_B_s_5 + 2*K_B_s_1 + K_B_s_2^2 + 2*K_B_s_2*K_B_s_3 + 2*K_B_s_2*K_B_s_4 + 2*K_B_s_2*K_B_s_5 + 2*K_B_s_2 + K_B_s_3^2 + 2*K_B_s_3*K_B_s_4 + 2*K_B_s_3*K_B_s_5 + 2*K_B_s_3 + K_B_s_4^2 + 2*K_B_s_4*K_B_s_5 + 2*K_B_s_4 + K_B_s_5^2 + 2*K_B_s_5 + 1)^(1/2) - K_A*Ltot + K_A*Rtot + 1)/(2*K_A) < Ltot and K_A^2*Ltot^2 + K_A^2*Rtot^2 + 2*K_A*K_B_s_1*Ltot + 2*K_A*K_B_s_1*Rtot + 2*K_A*K_B_s_2*Ltot + 2*K_A*K_B_s_2*Rtot + 2*K_A*K_B_s_3*Ltot + 2*K_A*K_B_s_3*Rtot + 2*K_A*K_B_s_4*Ltot + 2*K_A*K_B_s_4*Rtot + 2*K_A*K_B_s_5*Ltot + 2*K_A*K_B_s_5*Rtot + 2*K_A*Ltot + 2*K_A*Rtot + K_B_s_1^2 + 2*K_B_s_1*K_B_s_2 + 2*K_B_s_1*K_B_s_3 + 2*K_B_s_1*K_B_s_4 + 2*K_B_s_1*K_B_s_5 + 2*K_B_s_1 + K_B_s_2^2 + 2*K_B_s_2*K_B_s_3 + 2*K_B_s_2*K_B_s_4 + 2*K_B_s_2*K_B_s_5 + 2*K_B_s_2 + K_B_s_3^2 + 2*K_B_s_3*K_B_s_4 + 2*K_B_s_3*K_B_s_5 + 2*K_B_s_3 + K_B_s_4^2 + 2*K_B_s_4*K_B_s_5 + 2*K_B_s_4 + K_B_s_5^2 + 2*K_B_s_5 + 1 < 2*K_A^2*Ltot*Rtot and K_B_s_1 + K_B_s_2 + K_B_s_3 + K_B_s_4 + K_B_s_5 + (K_A^2*Ltot^2 - 2*K_A^2*Ltot*Rtot + K_A^2*Rtot^2 + 2*K_A*K_B_s_1*Ltot + 2*K_A*K_B_s_1*Rtot + 2*K_A*K_B_s_2*Ltot + 2*K_A*K_B_s_2*Rtot + 2*K_A*K_B_s_3*Ltot + 2*K_A*K_B_s_3*Rtot + 2*K_A*K_B_s_4*Ltot + 2*K_A*K_B_s_4*Rtot + 2*K_A*K_B_s_5*Ltot + 2*K_A*K_B_s_5*Rtot + 2*K_A*Ltot + 2*K_A*Rtot + K_B_s_1^2 + 2*K_B_s_1*K_B_s_2 + 2*K_B_s_1*K_B_s_3 + 2*K_B_s_1*K_B_s_4 + 2*K_B_s_1*K_B_s_5 + 2*K_B_s_1 + K_B_s_2^2 + 2*K_B_s_2*K_B_s_3 + 2*K_B_s_2*K_B_s_4 + 2*K_B_s_2*K_B_s_5 + 2*K_B_s_2 + K_B_s_3^2 + 2*K_B_s_3*K_B_s_4 + 2*K_B_s_3*K_B_s_5 + 2*K_B_s_3 + K_B_s_4^2 + 2*K_B_s_4*K_B_s_5 + 2*K_B_s_4 + K_B_s_5^2 + 2*K_B_s_5 + 1)^(1/2) + K_A*Rtot + 1 < K_A*Ltot, {}])](U_5R_derivation_images/math62.png)
Extract solutions:
solution_lines:=6:
eq3_19:= solution3_18[i,1] $ i=1..solution_lines;
nops(%)

![]()
Extract solutions
solution1:=eq3_19[1][1];
solution2:=eq3_19[2][1] ;
solution3:=eq3_19[4][1];
solution4:=eq3_19[5][1];




Analysis of solutions
If having a sequence of roots: Is 1st solution a combination of 2nd and 3rd?
/*
solutionA:=eq3_13[1]; // a sequence of roots
if solution2 in solutionA
then print(Unquoted,"First set of roots contains the second root.");
else print(Unquoted,"First set of roots DOES NOT contain the second root!");
end_if;
if solution3 in solutionA
then print(Unquoted,"First set of roots contains the third root.");
else print(Unquoted,"First set of roots DOES NOT contain the third root!");
end_if;
*/
Check correctness of the solution by substitution
eq3_18;
// Check the 1st solution
test1:=eq3_18 | Leq=solution1;
normal(%);
Simplify(%);
bool(%)
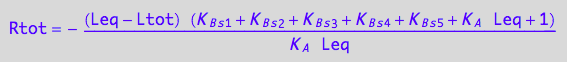


![]()
![]()
odd!
eq3_18;
// Check the 2nd solution
test1:=eq3_18 | Leq=solution2;
normal(%);
Simplify(%);
bool(%)
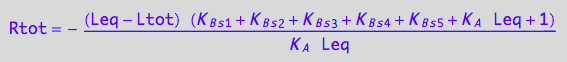


![]()
![]()
odd!
eq3_18;
// Check the 3rd solution
test1:=eq3_18 | Leq=solution3;
normal(%);
Simplify(%);
bool(%)
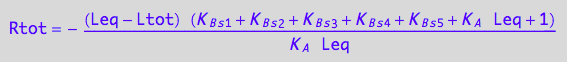



![]()
works...
eq3_18;
// Check the 4th solution
test1:=eq3_18 | Leq=solution4;
normal(%);
Simplify(%);
bool(%)
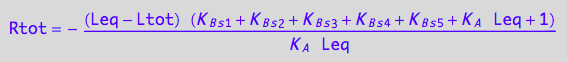



![]()
works...
=> two last solutions are confirmed by substitution. Proceed with these checking solutions numerically.
Test which solution is meaningful numerically
solution1;
% | K_A=1 | K_B_s_1=1 | K_B_s_2=1 | K_B_s_3=1 | K_B_s_4=1 | K_B_s_5=1 | Rtot=1 | Ltot=1;
float(%)

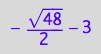
![]()
meaningless
solution2;
% | K_A=1 | K_B_s_1=1 | K_B_s_2=1 | K_B_s_3=1 | K_B_s_4=1 | K_B_s_5=1 | Rtot=1 | Ltot=1;
float(%)

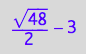
![]()
meaningful?
solution3;
% | K_A=1 | K_B_s_1=1 | K_B_s_2=1 | K_B_s_3=1 | K_B_s_4=1 | K_B_s_5=1 | Rtot=1 | Ltot=1;
float(%)

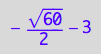
![]()
meaningless
solution4;
% | K_A=1 | K_B_s_1=1 | K_B_s_2=1 | K_B_s_3=1 | K_B_s_4=1 | K_B_s_5=1 | Rtot=1 | Ltot=1;
float(%)

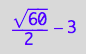
![]()
meaningful and different from solution 2!
Re-check positive solutions by substitution and calculation:
// Solution 2
test1:=eq3_18 | Leq=solution2;
result:= % | K_A=1 | K_B_s_1=1 | K_B_s_2=1 | K_B_s_3=1 | K_B_s_4=1 | K_B_s_5=1 | Rtot=1 | Ltot=1;
float(result);
Simplify(result);

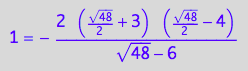
![]()
![]()
wrong solution!!!
Re-check by substitution and calculation:
// Solution 4
test1:=eq3_18 | Leq=solution4;
result:= % | K_A=1 | K_B_s_1=1 | K_B_s_2=1 | K_B_s_3=1 | K_B_s_4=1 | K_B_s_5=1 | Rtot=1 | Ltot=1;
float(result);
Simplify(result);

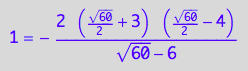
![]()
![]()
CORRECT!!!
Choose as a final solution
eq3_20:= Leq = solution4

Summary of equations for all species
Show species names
anames(Properties,User);
![]()
Name equations to recognize them in a different context
Leq_U_5R:=eq3_20

Req_U_5R:=eq3_17
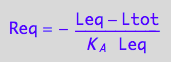
R_s_1eq_U_5R:=eq3_7
![]()
R_s_2eq_U_5R:=eq3_9
![]()
R_s_3eq_U_5R:=eq3_11
![]()
R_s_4eq_U_5R:=eq3_13
![]()
R_s_5eq_U_5R:=eq3_15
![]()
RLeq_U_5R:=eq3_1
![]()
Create functions for computing concentrations
Here I only check that the results are numerically meaningful--functions were created right. Scientific meaningfulness will be analyzed in a separate notebook 'Analysis'.
(use --> to force direct substitution):
fLeq_U_5R:= (Rtot, Ltot, K_A, K_B_s_1, K_B_s_2, K_B_s_3, K_B_s_4, K_B_s_5) --> Leq_U_5R[2]

//test operation
fLeq_U_5R(1,1,1,1,1,1,1,1):
float(%)
![]()
=> OK
fReq_U_5R:=(Rtot, Ltot, K_A, K_B_s_1, K_B_s_2, K_B_s_3, K_B_s_4, K_B_s_5) --> Req_U_5R[2] | Leq_U_5R

//test operation
fReq_U_5R(1,1,1,1,1,1,1,1):
float(%)
![]()
=> OK
fR_s_1eq_U_5R:=(Rtot, Ltot, K_A, K_B_s_1, K_B_s_2, K_B_s_3, K_B_s_4, K_B_s_5) --> R_s_1eq_U_5R[2] | Req_U_5R | Leq_U_5R

//test operation
fR_s_1eq_U_5R(1,1,1,1,1,1,1,1):
float(%)
![]()
=> OK
fR_s_2eq_U_5R:=(Rtot, Ltot, K_A, K_B_s_1, K_B_s_2, K_B_s_3, K_B_s_4, K_B_s_5) --> R_s_2eq_U_5R[2] | Req_U_5R | Leq_U_5R

//test operation
fR_s_2eq_U_5R(1,1,1,1,1,1,1,1):
float(%)
![]()
=> OK
fR_s_3eq_U_5R:=(Rtot, Ltot, K_A, K_B_s_1, K_B_s_2, K_B_s_3, K_B_s_4, K_B_s_5) --> R_s_3eq_U_5R[2] | Req_U_5R | Leq_U_5R

//test operation
fR_s_3eq_U_5R(1,1,1,1,1,1,1,1):
float(%)
![]()
=> OK
fR_s_4eq_U_5R:=(Rtot, Ltot, K_A, K_B_s_1, K_B_s_2, K_B_s_3, K_B_s_4, K_B_s_5) --> R_s_4eq_U_5R[2] | Req_U_5R | Leq_U_5R

//test operation
fR_s_4eq_U_5R(1,1,1,1,1,1,1,1):
float(%)
![]()
=> OK
fR_s_5eq_U_5R:=(Rtot, Ltot, K_A, K_B_s_1, K_B_s_2, K_B_s_3, K_B_s_4, K_B_s_5) --> R_s_5eq_U_5R[2] | Req_U_5R | Leq_U_5R

//test operation
fR_s_5eq_U_5R(1,1,1,1,1,1,1,1):
float(%)
![]()
=> OK
fRLeq_U_5R:=(Rtot, Ltot, K_A, K_B_s_1, K_B_s_2, K_B_s_3, K_B_s_4, K_B_s_5) --> RLeq_U_5R[2] | Req_U_5R | Leq_U_5R

//test operation
fRLeq_U_5R(1,1,1,1,1,1,1,1):
float(%)
![]()
=> OK
Check conservation laws (check the Rtot and Ltot are set to 1) :
fRLeq_U_5R(1,1,1,1,1,1,1,1)+fLeq_U_5R(1,1,1,1,1,1,1,1)
![]()
good
fRLeq_U_5R(1,1,1,1,1,1,1,1) + fReq_U_5R(1,1,1,1,1,1,1,1) + fR_s_1eq_U_5R(1,1,1,1,1,1,1,1) + fR_s_2eq_U_5R(1,1,1,1,1,1,1,1) + fR_s_3eq_U_5R(1,1,1,1,1,1,1,1) + fR_s_4eq_U_5R(1,1,1,1,1,1,1,1) + fR_s_5eq_U_5R(1,1,1,1,1,1,1,1) ;
float(%)
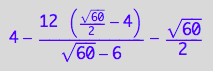
![]()
good!
(you can retrieve them later by executing: fread(filename,Quiet))
ProjectName
![]()
filename:=CurrentPath."/".ProjectName.".mb";
write(filename,
// Equations
Leq_U_5R,
Req_U_5R,
R_s_1eq_U_5R,
R_s_2eq_U_5R,
R_s_3eq_U_5R,
R_s_4eq_U_5R,
R_s_5eq_U_5R,
RLeq_U_5R,
// Analytical functions
fLeq_U_5R,
fReq_U_5R,
fR_s_1eq_U_5R,
fR_s_2eq_U_5R,
fR_s_3eq_U_5R,
fR_s_4eq_U_5R,
fR_s_5eq_U_5R,
fRLeq_U_5R
)
![]()
Conclusions
1. Analytical solution obtained.
2. Functions for analysis of behavior of solutions are created and saved.