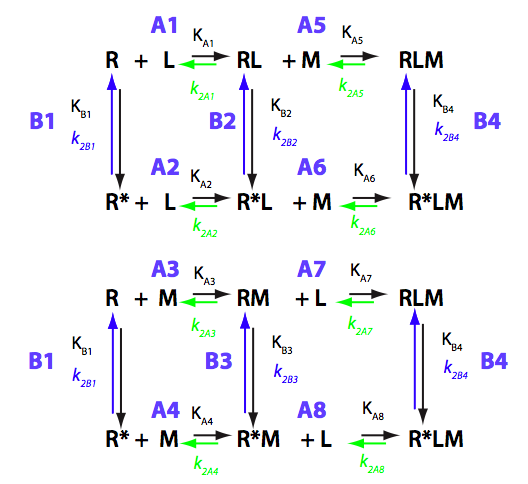
Equilibrium molar fractions of species
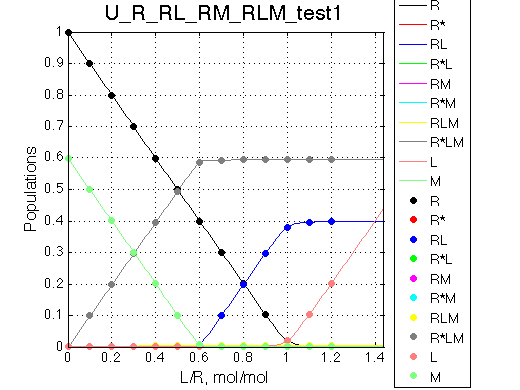
Other graphic formats: EPS, MATLAB Figure
Points correspond to steps in titration series corresponding to Rtotal and LRratio used in NMR line shape simulations below. Curves are smooth functions simulated at constant Rtotal. Populations of multimeric species are shown per monomer. Population of free ligand is normalized to total receptor concentration. |
Isothermal Titration Calorimetry profile
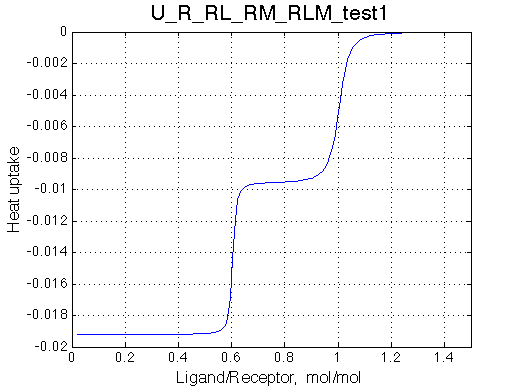
Other graphic formats: EPS, MATLAB Figure
Heat uptake curve is simulated as a derivative of the population of species multiplied by specific dH. |
U_R_RL_RM_RLM_test1
R* does not exist, L only binds to R, M does not bind to R, only to RL and RLM interconverts into R*LM
Equilibrium constants:
Ka(A1)=1.00e+06
Ka(A3)=1.00e-03
Ka(A5)=1.00e+05
Ka(B1)=1.00e-03
Ka(B2)=1.00e-03
Ka(B3)=1.00e-03
Ka(B4)=1.00e+02
Ka(A2)=1.00e+06 (dependent)
Ka(A4)=1.00e-03 (dependent)
Ka(A6)=1.00e+10 (dependent)
Ka(A7)=1.00e+14 (dependent)
Ka(A8)=1.00e+19 (dependent)
Rate constants (1-forward, 2-reverse):
k1(A1)=1.00e+06
k2(A1)=1.00e+00
k1(A2)=1.00e+06
k2(A2)=1.00e+00
k1(A3)=0.00e+00
k2(A3)=0.00e+00
k1(A4)=0.00e+00
k2(A4)=0.00e+00
k1(A5)=1.00e+05
k2(A5)=1.00e+00
k1(A6)=1.00e+10
k2(A6)=1.00e+00
k1(A7)=0.00e+00
k2(A7)=0.00e+00
k1(A8)=0.00e+00
k2(A8)=0.00e+00
k1(B1)=0.00e+00
k2(B1)=0.00e+00
k1(B2)=0.00e+00
k2(B2)=0.00e+00
k1(B3)=0.00e+00
k2(B3)=0.00e+00
k1(B4)=1.00e+02
k2(B4)=1.00e+00
Chemical shifts:
w0(R)=50.0 /s (8.0 Hz)
w0(R*)=0.0 /s (0.0 Hz)
w0(RL)=100.0 /s (15.9 Hz)
w0(R*L)=150.0 /s (23.9 Hz)
w0(RM)=0.0 /s (0.0 Hz)
w0(R*M)=0.0 /s (0.0 Hz)
w0(RLM)=200.0 /s (31.8 Hz)
w0(R*LM)=300.0 /s (47.8 Hz)
Base relaxation rates:
R2(R)=10.0 /s
R2(R*)=10.0 /s
R2(RL)=10.0 /s
R2(R*L)=10.0 /s
R2(RM)=10.0 /s
R2(R*M)=10.0 /s
R2(RLM)=10.0 /s
R2(R*LM)=10.0 /s
Enthalpy difference from the base state:
dH(R)=0.0
dH(R*)=0.0
dH(RL)=-1.0
dH(R*L)=-1.0
dH(RM)=0.0
dH(R*M)=0.0
dH(RLM)=-2.0
dH(R*LM)=-2.0
Total R concentration (*1000):
1.00 1.00 1.00 1.00 1.00 1.00 1.00 1.00 1.00 1.00 1.00 1.00 1.00
Ratio of total L to total R:
0.00 0.10 0.20 0.30 0.40 0.50 0.60 0.70 0.80 0.90 1.00 1.10 1.20 |
1D-spectra of the titration series
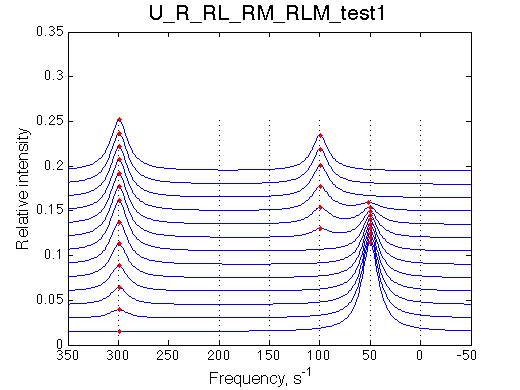
Other graphic formats: EPS, MATLAB Figure
Traces (bottom to top) correspond to L/R ratios indicated by points in the Populations graph (above). Red dots indicate peak maxima for a visual aid (also plotted as the chemical shift titration curves on the right). Dashed lines indicate chemical shifts of individual species. |
Chemical shift titration curves
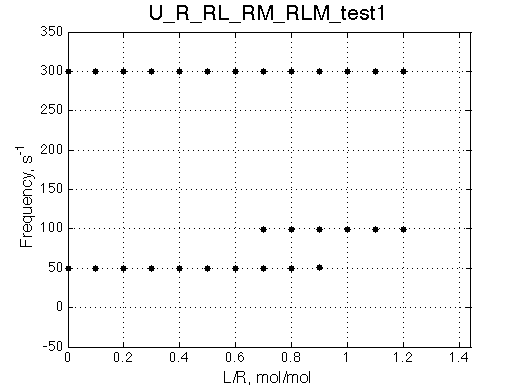
Other graphic formats: EPS, MATLAB Figure
Points correspond to positions of peak maxima numerically detected in the spectral traces. Sometimes not all peaks will be detected and signal/noise here is not considered. |
| Plot of the line widths (full width at a half height)
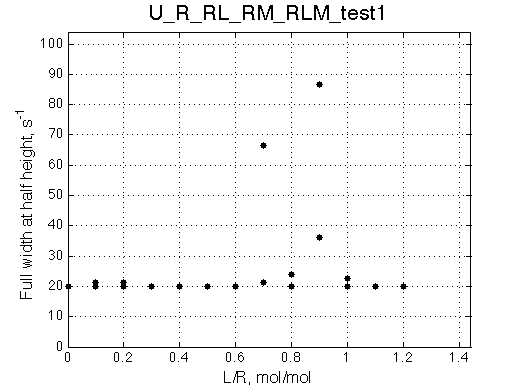
Plain text data for comparison plotting: TXT
Other graphic formats: EPS, MATLAB Figure
The full width at a half height (FWHH) is determined numerically for the detected peaks. For the lorentzian line shape FWHH = 2*R2.
At the points of peak coalescence the measurement fails, you should discard these points. The plain text data for re-plotting the graph is saved with the graphics. |
|
|